flowchart LR
S["<b>S</b><br/>Susceptible"] -->|"infection<br/>(se.flow)"| E
E["<b>E</b><br/>Exposed"] -->|"progression<br/>(ei.flow)"| I
I["<b>I</b><br/>Infectious"] -->|"recovery<br/>(ir.flow)"| R
R["<b>R</b><br/>Recovered"] -.->|"waning immunity<br/>(rs.flow, SEIRS only)"| S
style S fill:#3498db,color:#fff
style E fill:#f39c12,color:#fff
style I fill:#e74c3c,color:#fff
style R fill:#27ae60,color:#fff
SEIR/SEIRS Model: Adding an Exposed State
Overview
Many infectious diseases have an incubation period during which a person has been infected but is not yet infectious to others. Influenza, measles, and COVID-19 all exhibit this latent stage. The standard SIR model skips it entirely — newly infected individuals become immediately infectious — but adding an Exposed (E) compartment creates an SEIR model that captures this biologically important delay between infection and infectiousness.
This example shows how to extend EpiModel’s built-in SIR framework by replacing the infection and recovery modules with two custom functions: one that routes newly infected individuals into the E compartment instead of directly to I, and a progression module that handles the E-to-I-to-R cascade. The same progression module also supports an SEIRS extension where recovered individuals eventually lose immunity and return to the susceptible pool. This single parameter change — setting rs.rate above zero — fundamentally alters the long-run dynamics, transforming an epidemic that burns out into one that persists endemically.
The techniques demonstrated here — replacing a built-in infection module, writing a custom progression module, and tracking new compartment counts — are the building blocks for all the more complex models in the Gallery.
- model.R — Main simulation script
- module-fx.R — Custom module functions
Model Structure
| Compartment | Label | Description |
|---|---|---|
| Susceptible | S | Not infected; at risk of exposure |
| Exposed | E | Infected but not yet infectious (latent period) |
| Infectious | I | Infected and capable of transmitting to contacts |
| Recovered | R | Immune after recovery; permanent (SEIR) or temporary (SEIRS) |
Transmission occurs along discordant edges — partnerships where one node is susceptible and the other is infectious. For each discordant pair, the per-timestep transmission probability accounts for multiple acts per partnership: finalProb = 1 - (1 - inf.prob) ^ act.rate. With the default parameters (inf.prob = 0.5, act.rate = 2), this gives a per-partnership probability of 0.75 per timestep.
Each transition (E to I, I to R, R to S) is modeled as an independent Bernoulli trial at each timestep: every eligible individual has a constant probability of transitioning, regardless of how long they have been in their current compartment. This produces geometrically distributed waiting times with mean 1/rate.
Setup
suppressMessages(library(EpiModel))
nsims <- 5
ncores <- 5
nsteps <- 800Custom Modules
Infection Module
This module replaces EpiModel’s built-in infection function. The critical difference from a standard SIR infection module is that newly infected individuals enter the "e" (Exposed) state rather than becoming immediately infectious. The module extracts discordant edges via discord_edgelist(), computes per-partnership transmission probabilities accounting for multiple acts, and draws stochastic transmission events.
infect <- function(dat, at) {
## Attributes ##
active <- get_attr(dat, "active")
status <- get_attr(dat, "status")
infTime <- get_attr(dat, "infTime")
## Parameters ##
inf.prob <- get_param(dat, "inf.prob")
act.rate <- get_param(dat, "act.rate")
## Find infected nodes ##
idsInf <- which(active == 1 & status == "i")
nActive <- sum(active == 1)
nElig <- length(idsInf)
## Initialize default incidence at 0 ##
nInf <- 0
## If any infected nodes, proceed with transmission ##
if (nElig > 0 && nElig < nActive) {
## Look up discordant edgelist ##
1 del <- discord_edgelist(dat, at)
## If any discordant pairs, proceed ##
if (!(is.null(del))) {
# Set parameters on discordant edgelist data frame
del$transProb <- inf.prob
del$actRate <- act.rate
2 del$finalProb <- 1 - (1 - del$transProb)^del$actRate
# Stochastic transmission process
3 transmit <- rbinom(nrow(del), 1, del$finalProb)
# Keep rows where transmission occurred
del <- del[which(transmit == 1), ]
# Look up new ids if any transmissions occurred
idsNewInf <- unique(del$sus)
nInf <- length(idsNewInf)
# Set new attributes for those newly infected
if (nInf > 0) {
4 status[idsNewInf] <- "e"
infTime[idsNewInf] <- at
dat <- set_attr(dat, "status", status)
dat <- set_attr(dat, "infTime", infTime)
}
}
}
## Save summary statistic for S->E flow
dat <- set_epi(dat, "se.flow", at, nInf)
return(dat)
}- 1
-
discord_edgelist()returns all active edges where one partner is susceptible and the other infectious. - 2
-
Per-partnership probability: with
act.rateacts, the chance of at least one successful transmission is1 - P(all acts fail). - 3
- Each discordant pair independently transmits (or not) via a Bernoulli draw.
- 4
-
The key SEIR difference: newly infected individuals enter status
"e"(Exposed), not"i"(Infectious).
Progression Module
This module replaces EpiModel’s built-in recovery function and handles up to three transitions: E to I, I to R, and optionally R to S. The R-to-S waning immunity block only activates when rs.rate > 0, so a single function handles both SEIR and SEIRS dynamics without code duplication.
progress <- function(dat, at) {
## Attributes ##
active <- get_attr(dat, "active")
status <- get_attr(dat, "status")
## Parameters ##
ei.rate <- get_param(dat, "ei.rate")
ir.rate <- get_param(dat, "ir.rate")
rs.rate <- get_param(dat, "rs.rate")
## E to I progression process ##
nInf <- 0
idsEligInf <- which(active == 1 & status == "e")
nEligInf <- length(idsEligInf)
if (nEligInf > 0) {
1 vecInf <- which(rbinom(nEligInf, 1, ei.rate) == 1)
if (length(vecInf) > 0) {
idsInf <- idsEligInf[vecInf]
nInf <- length(idsInf)
status[idsInf] <- "i"
}
}
## I to R progression process ##
nRec <- 0
idsEligRec <- which(active == 1 & status == "i")
nEligRec <- length(idsEligRec)
if (nEligRec > 0) {
vecRec <- which(rbinom(nEligRec, 1, ir.rate) == 1)
if (length(vecRec) > 0) {
idsRec <- idsEligRec[vecRec]
nRec <- length(idsRec)
status[idsRec] <- "r"
}
}
## R to S waning immunity process (active only when rs.rate > 0) ##
nSus <- 0
2 if (rs.rate > 0) {
idsEligSus <- which(active == 1 & status == "r")
nEligSus <- length(idsEligSus)
if (nEligSus > 0) {
vecSus <- which(rbinom(nEligSus, 1, rs.rate) == 1)
if (length(vecSus) > 0) {
idsSus <- idsEligSus[vecSus]
nSus <- length(idsSus)
status[idsSus] <- "s"
}
}
}
## Write out updated status attribute ##
dat <- set_attr(dat, "status", status)
## Save summary statistics ##
dat <- set_epi(dat, "ei.flow", at, nInf)
dat <- set_epi(dat, "ir.flow", at, nRec)
dat <- set_epi(dat, "rs.flow", at, nSus)
3 dat <- set_epi(dat, "e.num", at, sum(active == 1 & status == "e"))
dat <- set_epi(dat, "r.num", at, sum(active == 1 & status == "r"))
return(dat)
}- 1
-
Each exposed individual independently transitions to infectious with probability
ei.rateper timestep (Bernoulli trial), producing geometrically distributed latent durations with mean1/ei.rate. - 2
-
The R-to-S block is skipped entirely when
rs.rate = 0, giving standard SEIR dynamics. Settingrs.rate > 0enables the SEIRS waning-immunity extension. - 3
-
Custom compartment counts (
e.num,r.num) must be tracked manually because the built-inprevalence.netmodule only knows abouts.num,i.num, andnum.
Network Model
n <- 500
nw <- network_initialize(n)
# Formation model: edges + isolates
1formation <- ~edges + isolates
target.stats <- c(150, 240)
# Partnership duration: mean of 25 timesteps
2coef.diss <- dissolution_coefs(dissolution = ~offset(edges), duration = 25)
coef.diss
# Fit the ERGM
est <- netest(nw, formation, target.stats, coef.diss)- 1
- With 150 edges (mean degree = 0.6) and 240 isolates (48% of nodes with no active partnership), this produces a sparse network where roughly half the population has no current contact.
- 2
-
Closed population (no vital dynamics), so no mortality adjustment (
d.rate) is needed for dissolution coefficients.
Dissolution Coefficients
=======================
Dissolution Model: ~offset(edges)
Target Statistics: 25
Crude Coefficient: 3.178054
Mortality/Exit Rate: 0
Adjusted Coefficient: 3.178054Network Diagnostics
dx <- netdx(est, nsims = nsims, ncores = ncores, nsteps = nsteps,
nwstats.formula = ~edges + isolates + degree(0:5))
Network Diagnostics
-----------------------
- Simulating 5 networks
- Calculating formation statisticsprint(dx)EpiModel Network Diagnostics
=======================
Diagnostic Method: Dynamic
Simulations: 5
Time Steps per Sim: 800
Formation Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 150 147.084 -1.944 0.706 -4.129 2.213 8.866
isolates 240 244.281 1.784 0.964 4.443 2.767 13.502
degree0 NA 244.281 NA 0.964 NA 2.767 13.502
degree1 NA 220.807 NA 0.819 NA 1.556 12.248
degree2 NA 31.594 NA 0.387 NA 1.422 5.714
degree3 NA 3.101 NA 0.112 NA 0.230 1.776
degree4 NA 0.212 NA 0.023 NA 0.049 0.456
degree5 NA 0.004 NA 0.002 NA 0.003 0.063
Duration Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 25 24.963 -0.148 0.185 -0.2 0.617 1.911
Dissolution Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.04 0.04 -0.428 0 -0.73 0.001 0.016plot(dx)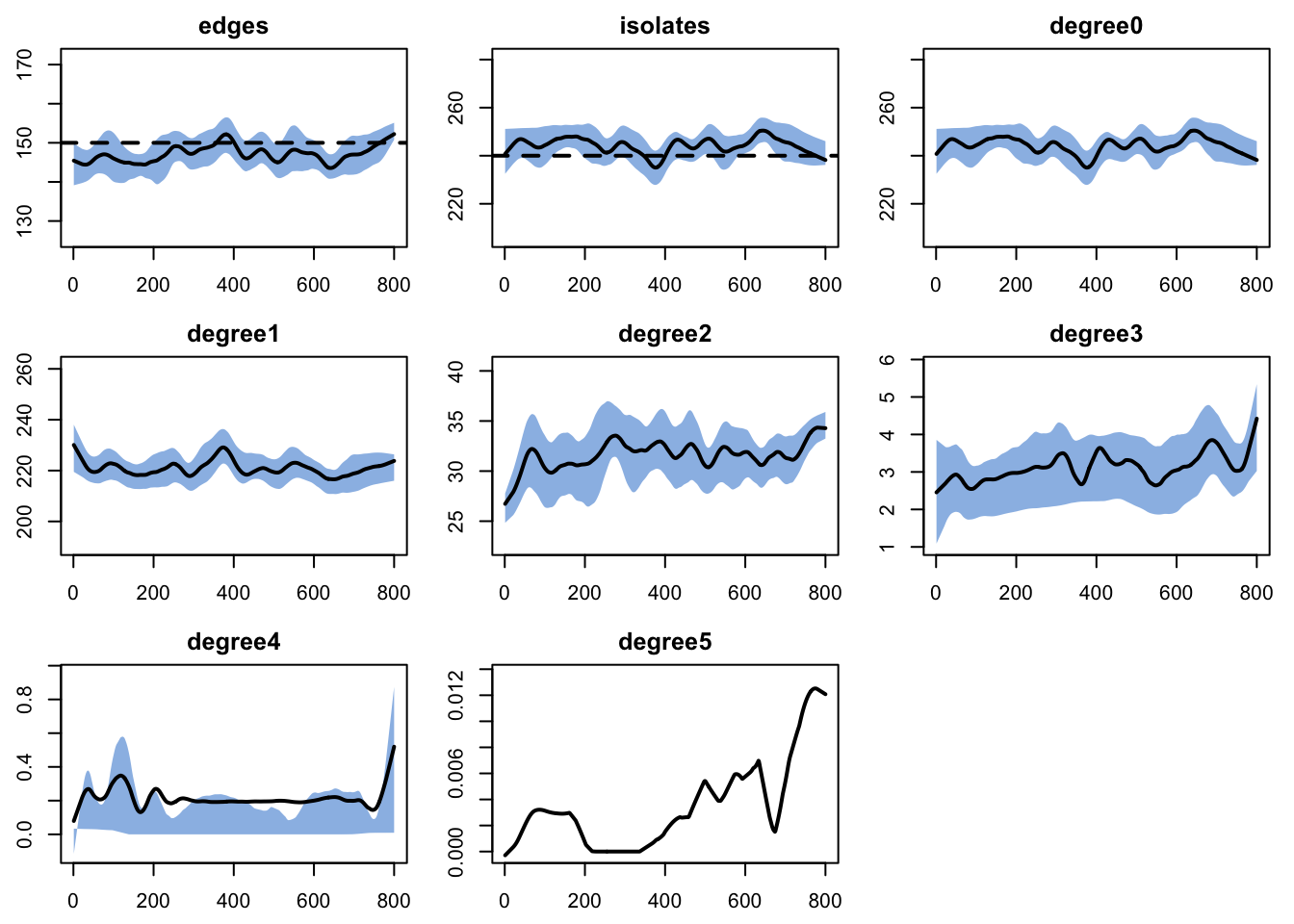
Epidemic Simulation
Initial Conditions
The epidemic is seeded with 10 infectious nodes (2% prevalence).
init <- init.net(i.num = 10)Scenario 1: SEIR (Permanent Immunity)
In the SEIR model, recovered individuals gain permanent immunity (rs.rate = 0). The epidemic burns through the susceptible population and dies out once the effective reproduction number drops below 1.
param_seir <- param.net(
inf.prob = 0.5,
act.rate = 2,
1 ei.rate = 1 / 50,
ir.rate = 1 / 75,
2 rs.rate = 0
)
control <- control.net(
3 type = NULL,
nsteps = nsteps,
nsims = nsims,
ncores = ncores,
infection.FUN = infect,
progress.FUN = progress
)
sim_seir <- netsim(est, param_seir, init, control)
print(sim_seir)- 1
-
Mean latent (E) duration of 50 timesteps; mean infectious (I) duration of 75 timesteps. These are stylized for pedagogical clarity — for influenza, typical values would be
ei.rate ~ 1/2andir.rate ~ 1/5. - 2
-
rs.rate = 0means permanent immunity after recovery (standard SEIR). - 3
-
type = NULLsignals that all disease modules are custom-supplied, not built-in.
EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type:
No. simulations: 5
No. time steps: 800
No. NW groups: 1
Fixed Parameters
---------------------------
inf.prob = 0.5
act.rate = 2
ei.rate = 0.02
ir.rate = 0.01333333
rs.rate = 0
groups = 1
Model Functions
-----------------------
initialize.FUN
resim_nets.FUN
summary_nets.FUN
infection.FUN
nwupdate.FUN
prevalence.FUN
verbose.FUN
progress.FUN
Model Output
-----------------------
Variables: s.num i.num num ei.flow ir.flow rs.flow e.num
r.num se.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 150 147.382 -1.745 0.674 -3.885 1.746 9.043
isolates 240 244.291 1.788 1.025 4.188 2.430 13.906
Duration Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 25 25.016 0.065 0.223 0.073 0.52 2.119
Dissolution Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.04 0.04 1.031 0 1.585 0.001 0.016Scenario 2: SEIRS (Waning Immunity)
Adding waning immunity (rs.rate = 1/100) fundamentally changes the long-run dynamics. Instead of epidemic burnout, the disease persists indefinitely as recovered individuals return to the susceptible pool, creating sustained endemic transmission with periodic fluctuations.
param_seirs <- param.net(
inf.prob = 0.5,
act.rate = 2,
ei.rate = 1 / 50,
ir.rate = 1 / 75,
1 rs.rate = 1 / 100
)
sim_seirs <- netsim(est, param_seirs, init, control)
print(sim_seirs)- 1
- Mean immune duration of 100 timesteps before returning to S. This single change transforms the dynamics from epidemic burnout to endemic persistence.
EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type:
No. simulations: 5
No. time steps: 800
No. NW groups: 1
Fixed Parameters
---------------------------
inf.prob = 0.5
act.rate = 2
ei.rate = 0.02
ir.rate = 0.01333333
rs.rate = 0.01
groups = 1
Model Functions
-----------------------
initialize.FUN
resim_nets.FUN
summary_nets.FUN
infection.FUN
nwupdate.FUN
prevalence.FUN
verbose.FUN
progress.FUN
Model Output
-----------------------
Variables: s.num i.num num ei.flow ir.flow rs.flow e.num
r.num se.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 150 146.974 -2.017 0.752 -4.025 1.817 9.231
isolates 240 244.928 2.053 0.959 5.138 2.162 13.341
Duration Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 25 24.802 -0.791 0.2 -0.987 0.445 1.933
Dissolution Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.04 0.04 1.116 0 1.709 0 0.017Analysis
sim_seir <- mutate_epi(sim_seir, prev = i.num / num)
sim_seirs <- mutate_epi(sim_seirs, prev = i.num / num)Prevalence Comparison
The central question: how does waning immunity change disease burden? In the SEIR model, prevalence rises then falls to zero as immunity accumulates and susceptibles are exhausted. In the SEIRS model, prevalence stabilizes at an endemic level because recovered individuals continuously return to the susceptible pool.
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_seir, y = "prev",
main = "Prevalence: SEIR vs. SEIRS",
ylab = "Prevalence (I / N)", xlab = "Time Step",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim_seirs, y = "prev",
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE, add = TRUE)
legend("topright", legend = c("SEIR", "SEIRS"),
col = c("firebrick", "steelblue"), lwd = 2, bty = "n")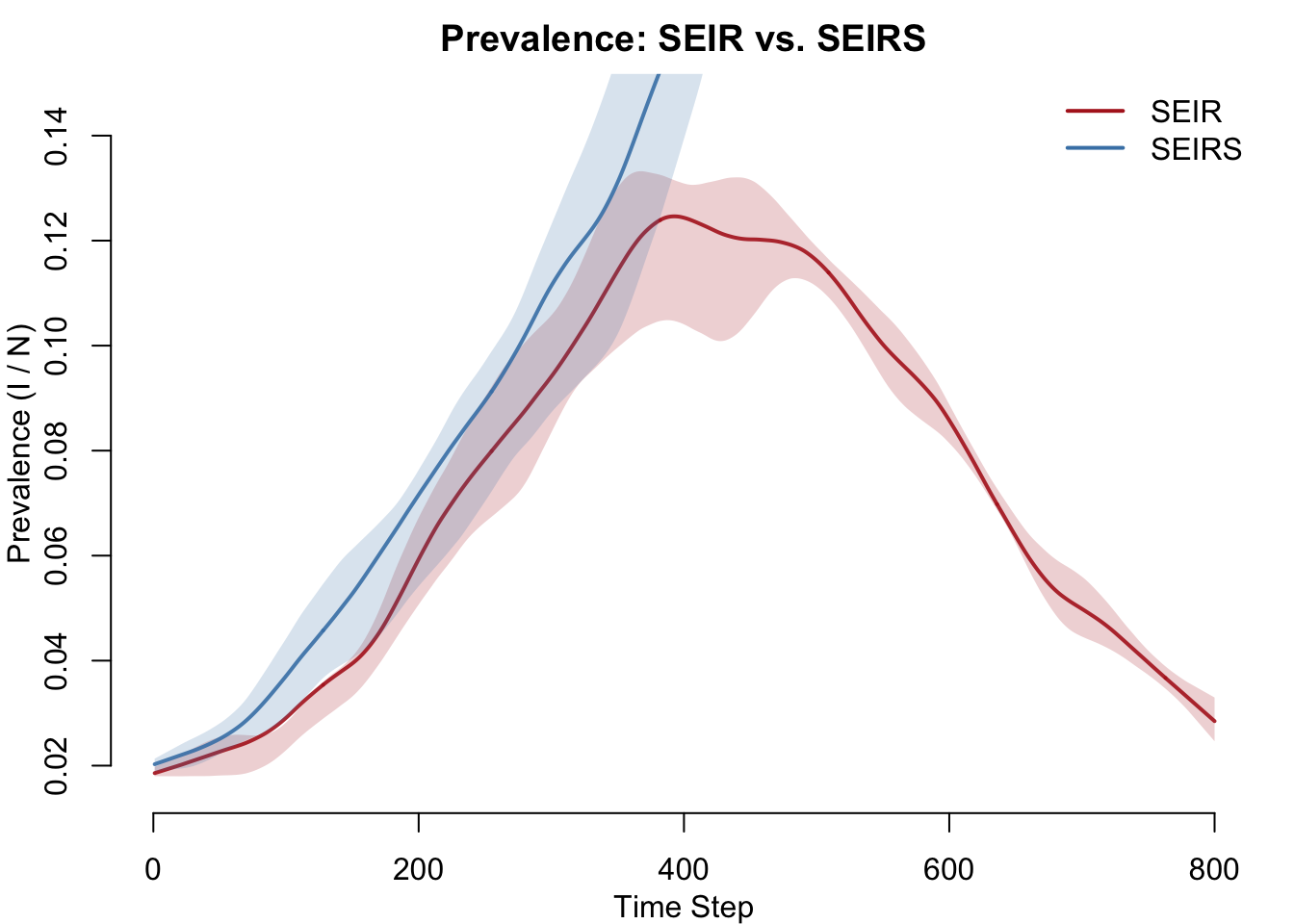
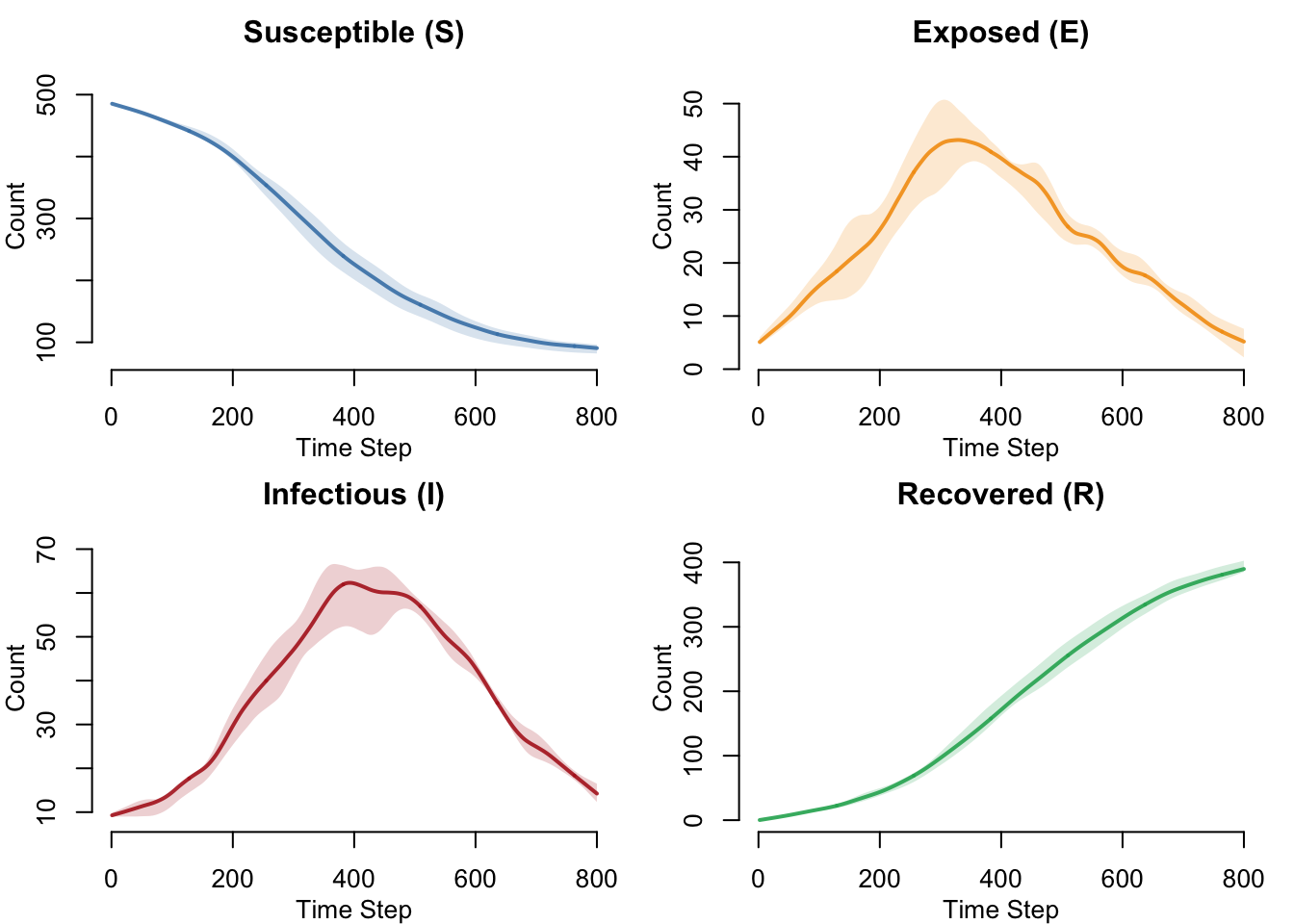
SEIR Compartment Dynamics
All four compartments over time in the standard SEIR model. The classic SEIR trajectory shows S declining as individuals are infected, E and I rising then falling as the epidemic wave passes, and R accumulating toward the full population as immunity builds.
compartments <- c("s.num", "e.num", "i.num", "r.num")
titles <- c("Susceptible (S)", "Exposed (E)", "Infectious (I)", "Recovered (R)")
colors <- c("steelblue", "#f39c12", "firebrick", "#27ae60")
par(mfrow = c(2, 2), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
for (j in seq_along(compartments)) {
plot(sim_seir, y = compartments[j],
main = titles[j], ylab = "Count", xlab = "Time Step",
mean.col = colors[j], mean.lwd = 2, mean.smooth = TRUE,
qnts.col = colors[j], qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
}
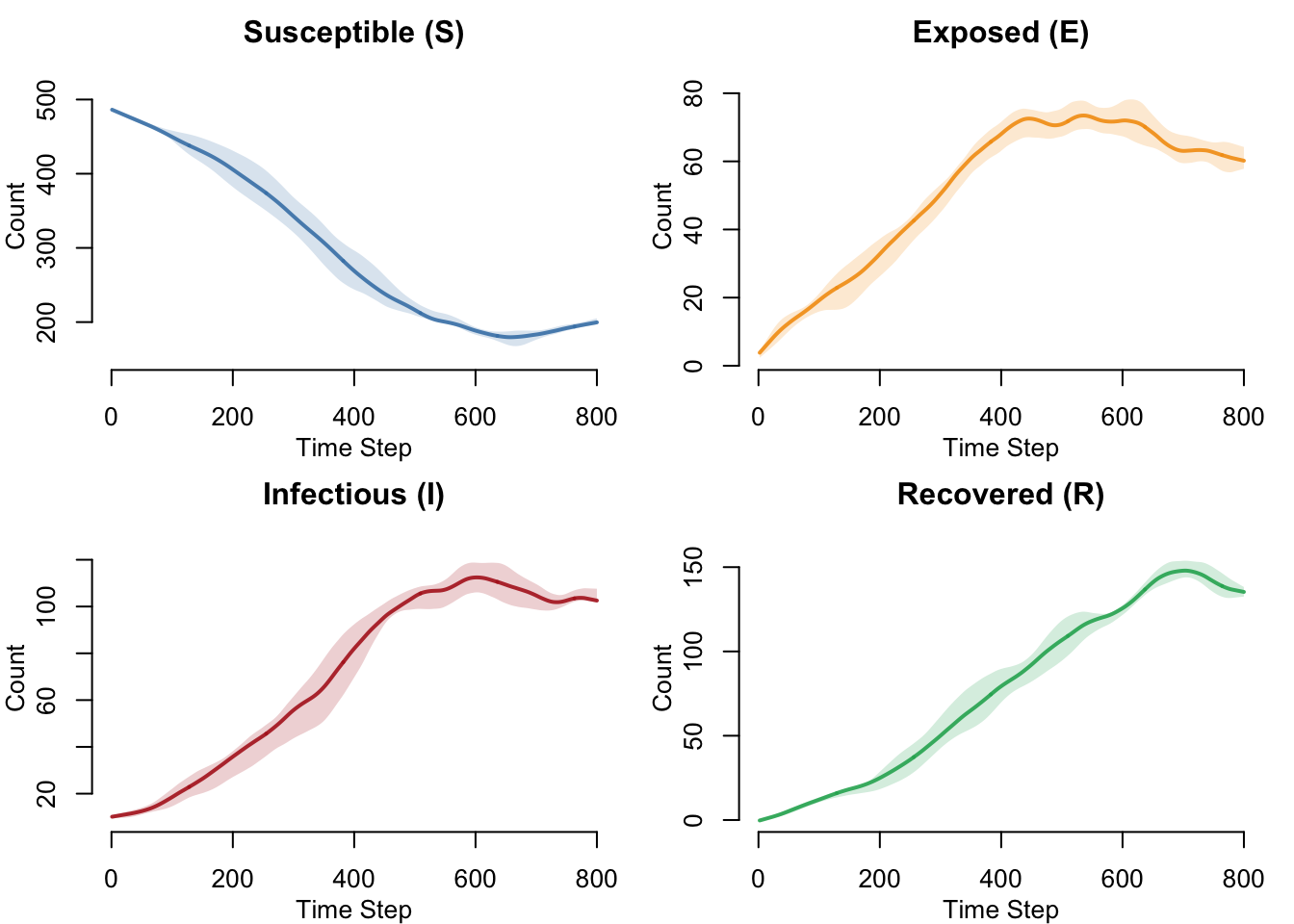
SEIRS Compartment Dynamics
With waning immunity, S does not deplete permanently and R does not accumulate forever. The system reaches a dynamic equilibrium with all four compartments maintaining nonzero populations — a hallmark of endemic disease dynamics.
par(mfrow = c(2, 2), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
for (j in seq_along(compartments)) {
plot(sim_seirs, y = compartments[j],
main = titles[j], ylab = "Count", xlab = "Time Step",
mean.col = colors[j], mean.lwd = 2, mean.smooth = TRUE,
qnts.col = colors[j], qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
}
New Infections (Incidence)
In the SEIR model, incidence peaks then declines to zero as susceptibles are exhausted. In the SEIRS model, incidence is sustained by the continuous resupply of susceptibles from recovered individuals losing immunity.
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_seir, y = "se.flow",
main = "New Infections: SEIR vs. SEIRS",
ylab = "New Infections per Timestep", xlab = "Time Step",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim_seirs, y = "se.flow",
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE, add = TRUE)
legend("topright", legend = c("SEIR", "SEIRS"),
col = c("firebrick", "steelblue"), lwd = 2, bty = "n")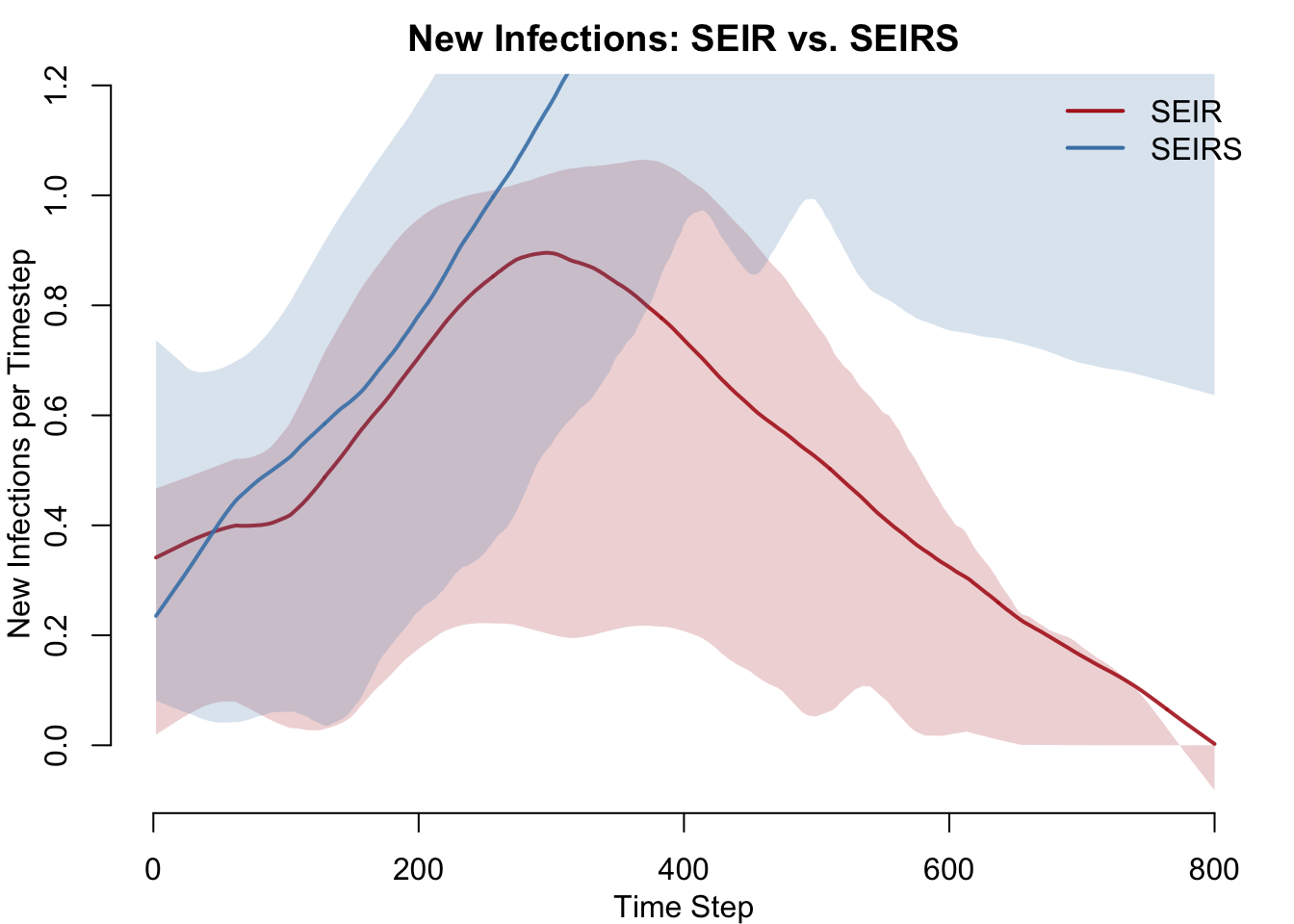
Summary Table
df_seir <- as.data.frame(sim_seir)
df_seirs <- as.data.frame(sim_seirs)
knitr::kable(data.frame(
Metric = c("Peak prevalence",
"Mean prevalence",
"Cumulative infections",
"Final recovered count"),
SEIR = c(
round(max(tapply(df_seir$prev, df_seir$sim, max, na.rm = TRUE)), 3),
round(mean(df_seir$prev, na.rm = TRUE), 3),
round(mean(tapply(df_seir$se.flow, df_seir$sim, sum, na.rm = TRUE))),
round(mean(df_seir$r.num[df_seir$time == max(df_seir$time)], na.rm = TRUE))
),
SEIRS = c(
round(max(tapply(df_seirs$prev, df_seirs$sim, max, na.rm = TRUE)), 3),
round(mean(df_seirs$prev, na.rm = TRUE), 3),
round(mean(tapply(df_seirs$se.flow, df_seirs$sim, sum, na.rm = TRUE))),
round(mean(df_seirs$r.num[df_seirs$time == max(df_seirs$time)], na.rm = TRUE))
)
))| Metric | SEIR | SEIRS |
|---|---|---|
| Peak prevalence | 0.180 | 0.264 |
| Mean prevalence | 0.073 | 0.142 |
| Cumulative infections | 400.000 | 918.000 |
| Final recovered count | 390.000 | 137.000 |
Parameters
Transmission
| Parameter | Description | Default |
|---|---|---|
inf.prob |
Per-act transmission probability | 0.5 |
act.rate |
Acts per partnership per timestep | 2 |
Disease Progression
| Parameter | Description | Default |
|---|---|---|
ei.rate |
E-to-I rate (1 / mean latent duration) | 1/50 |
ir.rate |
I-to-R rate (1 / mean infectious duration) | 1/75 |
rs.rate |
R-to-S waning immunity rate (0 = permanent immunity) | 0 (SEIR) or 1/100 (SEIRS) |
Network
| Parameter | Description | Default |
|---|---|---|
| Population size | Number of nodes | 500 |
| Target edges | Mean concurrent partnerships | 150 |
| Target isolates | Nodes with degree 0 | 240 |
| Partnership duration | Mean edge duration (timesteps) | 25 |
These parameters are stylized for pedagogical clarity, not calibrated to a specific pathogen. For influenza, typical values would be ei.rate ~ 1/2 and ir.rate ~ 1/5.
Module Execution Order
resim_nets → infection (infect) → progress → prevalenceThe built-in prevalence.net module runs last and computes s.num, i.num, and num from the updated status attribute. The custom progress module additionally tracks e.num, r.num, and all transition flows (ei.flow, ir.flow, rs.flow).
Next Steps
- Add vital dynamics (births and deaths) to maintain population size over long runs and allow true endemic equilibrium — see SI with Vital Dynamics
- Incorporate vaccination using all-or-nothing or leaky mechanisms — see SEIR with AON Vaccination and SEIRS with Leaky Vaccination
- Vary progression rates by individual attributes (e.g., age, immune status) to model heterogeneous disease courses
- Add stage-dependent infectiousness where transmission probability depends on disease stage — see the HIV model for this pattern
- Calibrate to a specific pathogen by substituting published estimates for latent and infectious period durations