flowchart LR
S["<b>S</b><br/>Susceptible"] -->|"infection<br/>(si.flow)"| I["<b>I</b><br/>Infectious"]
style S fill:#3498db,color:#fff
style I fill:#e74c3c,color:#fff
Epidemics over Observed Networks
Overview
The standard EpiModel workflow begins with egocentrically observed network data, fits a temporal ERGM (exponential random graph model) with the generative network effects of interest, and then simulates from that model fit over time. This example demonstrates an alternative approach: using a dynamic network census where all nodes and edges are directly observed over a series of discrete time steps. Rather than fitting and simulating from a statistical network model, the observed networkDynamic object is placed directly into the simulation data structure.
This is the only gallery example that bypasses EpiModel’s ERGM estimation and simulation pipeline entirely. The resimulate.network parameter is set to FALSE so the network is used as-is rather than being regenerated at each timestep. Because there is no model fit, this approach requires custom initialization and infection modules that handle the observed network directly.
Two scenarios are demonstrated using a simulated dataset from the networkDynamicData package (treated as if it were an observed network census): a basic SI model with constant transmission probability, and a two-stage disease model with time-varying transmission probability and network visualization. An important caveat — the observed network has edge activity recorded over a finite window, so nsteps must match the number of observed time steps — is also explored.
Download the individual source files for this example:
model.R— Main simulation scriptmodule-fx.R— Custom module functions
Model Structure
Compartments
| Compartment | Label | Description |
|---|---|---|
| Susceptible | S | Not infected; at risk of infection through network contacts |
| Infectious | I | Infected; can transmit to susceptible partners |
Flow Diagram
This is a simple SI model with no recovery. In a closed population without vital dynamics, prevalence increases monotonically toward saturation.
Observed Network Data
Unlike other gallery examples, this model does not call netest() to fit an ERGM. Instead, the observed networkDynamic object contains edge spells (onset/terminus times for each partnership) recorded over approximately 100 discrete time steps. Key characteristics of the network:
- 1000 nodes in an undirected network
- Edge spells define when each partnership is active
- The existing
status.activevertex attribute is removed before simulation, since we simulate our own epidemic - No network resimulation — the observed edge structure is fixed throughout the simulation
Setup
library(EpiModel)
library(networkDynamicData)Simulation Parameters
nsims <- 5
ncores <- 5
nsteps <- 100Custom Modules
This example requires two custom modules because the standard EpiModel initialization and infection modules expect a netest object (a fitted ERGM). Since we are working with an observed network that has no model fit, we need custom functions that handle the network directly.
Initialization Module
The init_obsnw function replaces EpiModel’s built-in initialize.net. It creates the master data list, places the observed networkDynamic object directly into the simulation data structure, initializes node attributes, and randomly assigns initial infections. When track.nw.attr = TRUE, it also stores time-varying disease status on the network as a temporally extended attribute (TEA), enabling network visualization with color-coded disease status.
init_obsnw <- function(x, param, init, control, s) {
1 dat <- create_dat_object(param, init, control)
dat$num.nw <- 1L
2 dat$run$nw[[1]] <- x
dat <- set_param(dat, "groups", 1)
i.num <- get_init(dat, "i.num")
n <- network.size(dat$run$nw[[1]])
dat <- append_core_attr(dat, 1, n)
status <- rep("s", n)
3 status[sample(1:n, i.num)] <- "i"
dat <- set_attr(dat, "status", status)
infTime <- rep(NA, n)
infTime[which(status == "i")] <- 1
dat <- set_attr(dat, "infTime", infTime)
track.nw.attr <- get_param(dat, "track.nw.attr",
override.null.error = TRUE)
if (!is.null(track.nw.attr) && track.nw.attr) {
4 dat$run$nw[[1]] <- networkDynamic::activate.vertex.attribute(
dat$run$nw[[1]], prefix = "testatus",
value = get_attr(dat, "status"), onset = 1, terminus = Inf
)
}
5 dat <- prevalence.net(dat, 1)
return(dat)
}- 1
- Create the master data list from parameters, initial conditions, and control settings.
- 2
-
Place the observed
networkDynamicobject directly into the data structure — no ERGM fit needed. - 3
-
Randomly assign
i.numinitial infections among thennetwork nodes. - 4
-
When
track.nw.attr = TRUE, store disease status as a temporally extended attribute on the network for visualization. - 5
-
Call the built-in
prevalence.net()to record initial epidemic statistics.
Infection Module
The infect_obsnw function handles transmission over the observed network. It supports two modes selected automatically based on which parameters are provided: a simple mode with constant transmission probability (inf.prob) and a time-varying mode where transmission depends on infection duration (inf.prob.stage1 / inf.prob.stage2).
infect_obsnw <- function(dat, at) {
active <- get_attr(dat, "active")
status <- get_attr(dat, "status")
infTime <- get_attr(dat, "infTime")
act.rate <- get_param(dat, "act.rate")
inf.prob.stage1 <- get_param(dat, "inf.prob.stage1",
override.null.error = TRUE)
1 time_varying <- !is.null(inf.prob.stage1)
if (time_varying) {
inf.prob.stage2 <- get_param(dat, "inf.prob.stage2")
dur.stage1 <- get_param(dat, "dur.stage1")
} else {
inf.prob <- get_param(dat, "inf.prob")
}
idsInf <- which(active == 1 & status == "i")
nActive <- sum(active == 1)
nElig <- length(idsInf)
totInf <- 0
if (nElig > 0 && nElig < nActive) {
2 del <- discord_edgelist(dat, at)
if (!is.null(del)) {
if (time_varying) {
infDur.del <- at - infTime[del$inf]
3 del$transProb <- ifelse(infDur.del <= dur.stage1,
inf.prob.stage1, inf.prob.stage2)
} else {
del$transProb <- inf.prob
}
del$actRate <- act.rate
del$finalProb <- 1 - (1 - del$transProb) ^ del$actRate
transmit <- rbinom(nrow(del), 1, del$finalProb)
del <- del[which(transmit == 1), ]
idsNewInf <- unique(del$sus)
totInf <- length(idsNewInf)
if (totInf > 0) {
status[idsNewInf] <- "i"
infTime[idsNewInf] <- at
dat <- set_attr(dat, "status", status)
dat <- set_attr(dat, "infTime", infTime)
track.nw.attr <- get_param(dat, "track.nw.attr",
override.null.error = TRUE)
if (!is.null(track.nw.attr) && track.nw.attr) {
dat$run$nw[[1]] <-
4 networkDynamic::activate.vertex.attribute(
dat$run$nw[[1]], prefix = "testatus",
value = "i", onset = at, terminus = Inf, v = idsNewInf
)
}
}
}
}
5 dat <- set_epi(dat, "si.flow", at, totInf)
return(dat)
}- 1
-
Automatically detect whether to use time-varying transmission based on whether
inf.prob.stage1was specified in the parameters. - 2
-
Use
discord_edgelist()to identify susceptible–infectious partnerships active at the current timestep. - 3
-
In time-varying mode, assign transmission probability based on how long each infector has been infected relative to
dur.stage1. - 4
- Update the temporally extended attribute on the network so newly infected nodes are visualized correctly.
- 5
-
Record the number of new infections at this timestep as the
si.flowepidemic tracker.
Network Model
Loading the Observed Network
The concurrencyComparisonNets dataset from the networkDynamicData package provides an observed dynamic network. We extract the base network and remove the existing disease status attribute, since we will simulate our own epidemic.
data(concurrencyComparisonNets)
nw <- base
print(nw)NetworkDynamic properties:
distinct change times: 104
maximal time range: -Inf until Inf
Dynamic (TEA) attributes:
Vertex TEAs: status.active
Includes optional net.obs.period attribute:
Network observation period info:
Number of observation spells: 1
Maximal time range observed: 2 until 102
Temporal mode: discrete
Time unit: step
Suggested time increment: 1
Network attributes:
vertices = 1000
directed = FALSE
hyper = FALSE
loops = FALSE
multiple = FALSE
bipartite = FALSE
net.obs.period: (not shown)
total edges= 1916
missing edges= 0
non-missing edges= 1916
Vertex attribute names:
status.active vertex.names
Edge attribute names not shown nw <- network::delete.vertex.attribute(nw, "status.active")Epidemic Simulation
Example 1: Basic SI Epidemic
The first scenario uses a simple SI model with a constant per-act transmission probability of 50% and a single act per partnership per timestep.
param <- param.net(inf.prob = 0.5, act.rate = 1)
init <- init.net(i.num = 10)The control settings specify our custom modules and disable network resimulation and network statistics saving (since there is no ERGM fit to compute statistics from). The nsteps parameter is set to match the number of observed time steps in the network.
control <- control.net(
type = NULL,
nsteps = nsteps,
nsims = nsims,
ncores = ncores,
initialize.FUN = init_obsnw,
infection.FUN = infect_obsnw,
prevalence.FUN = prevalence.net,
resimulate.network = FALSE,
save.nwstats = FALSE,
verbose = FALSE
)sim <- netsim(nw, param, init, control)
print(sim)EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type:
No. simulations: 5
No. time steps: 100
No. NW groups: 1
Fixed Parameters
---------------------------
inf.prob = 0.5
act.rate = 1
groups = 1
Model Functions
-----------------------
initialize.FUN
resim_nets.FUN
summary_nets.FUN
infection.FUN
nwupdate.FUN
prevalence.FUN
verbose.FUN
Model Output
-----------------------
Variables: s.num i.num num si.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5df <- as.data.frame(sim)
head(df, 25) sim time s.num i.num num si.flow
1 1 1 990 10 1000 NA
2 1 2 987 13 1000 3
3 1 3 982 18 1000 5
4 1 4 976 24 1000 6
5 1 5 974 26 1000 2
6 1 6 967 33 1000 7
7 1 7 966 34 1000 1
8 1 8 964 36 1000 2
9 1 9 961 39 1000 3
10 1 10 959 41 1000 2
11 1 11 957 43 1000 2
12 1 12 955 45 1000 2
13 1 13 951 49 1000 4
14 1 14 946 54 1000 5
15 1 15 945 55 1000 1
16 1 16 943 57 1000 2
17 1 17 939 61 1000 4
18 1 18 935 65 1000 4
19 1 19 932 68 1000 3
20 1 20 930 70 1000 2
21 1 21 927 73 1000 3
22 1 22 925 75 1000 2
23 1 23 920 80 1000 5
24 1 24 920 80 1000 0
25 1 25 916 84 1000 4Caveats: Simulating Past the Observation Window
The observed network has edge activity recorded over a finite window of approximately 100 timesteps. Edges that are active at the last observed time remain active indefinitely (a networkDynamic convention). This means nothing prevents simulating past the observation window — but the results become meaningless: the frozen edge set no longer reflects real dynamics, prevalence plateaus, and incidence drops to zero.
head(get.dyads.active(nw, at = 1), 10) [,1] [,2]
[1,] 834 4
[2,] 251 9
[3,] 722 9
[4,] 93 10
[5,] 169 10
[6,] 37 12
[7,] 309 13
[8,] 295 23
[9,] 187 26
[10,] 564 27head(get.dyads.active(nw, at = 100), 10) [,1] [,2]
[1,] 445 46
[2,] 464 78
[3,] 380 270
[4,] 777 365
[5,] 850 808
[6,] 14 852
[7,] 308 868
[8,] 664 711
[9,] 428 930
[10,] 534 780head(get.dyads.active(nw, at = 200), 10) [,1] [,2]
[1,] 445 46
[2,] 464 78
[3,] 380 270
[4,] 777 365
[5,] 850 808
[6,] 14 852
[7,] 664 711
[8,] 428 930
[9,] 534 780
[10,] 50 242To demonstrate this problem, we simulate 200 timesteps on the approximately 100-step network:
control_caveat <- control.net(
type = NULL,
nsteps = 200,
nsims = nsims,
ncores = ncores,
initialize.FUN = init_obsnw,
infection.FUN = infect_obsnw,
prevalence.FUN = prevalence.net,
resimulate.network = FALSE,
save.nwstats = FALSE,
verbose = FALSE
)
sim_caveat <- netsim(nw, param, init, control_caveat)par(mfrow = c(1, 2), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_caveat, y = "i.num",
main = "Prevalence (Past Observations)",
ylab = "Infected Count", xlab = "Time Step",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = FALSE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = FALSE,
legend = FALSE)
plot(sim_caveat, y = "si.flow",
main = "Incidence (Past Observations)",
ylab = "New Infections", xlab = "Time Step",
mean.col = "steelblue", mean.lwd = 2, mean.smooth = FALSE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = FALSE,
legend = FALSE)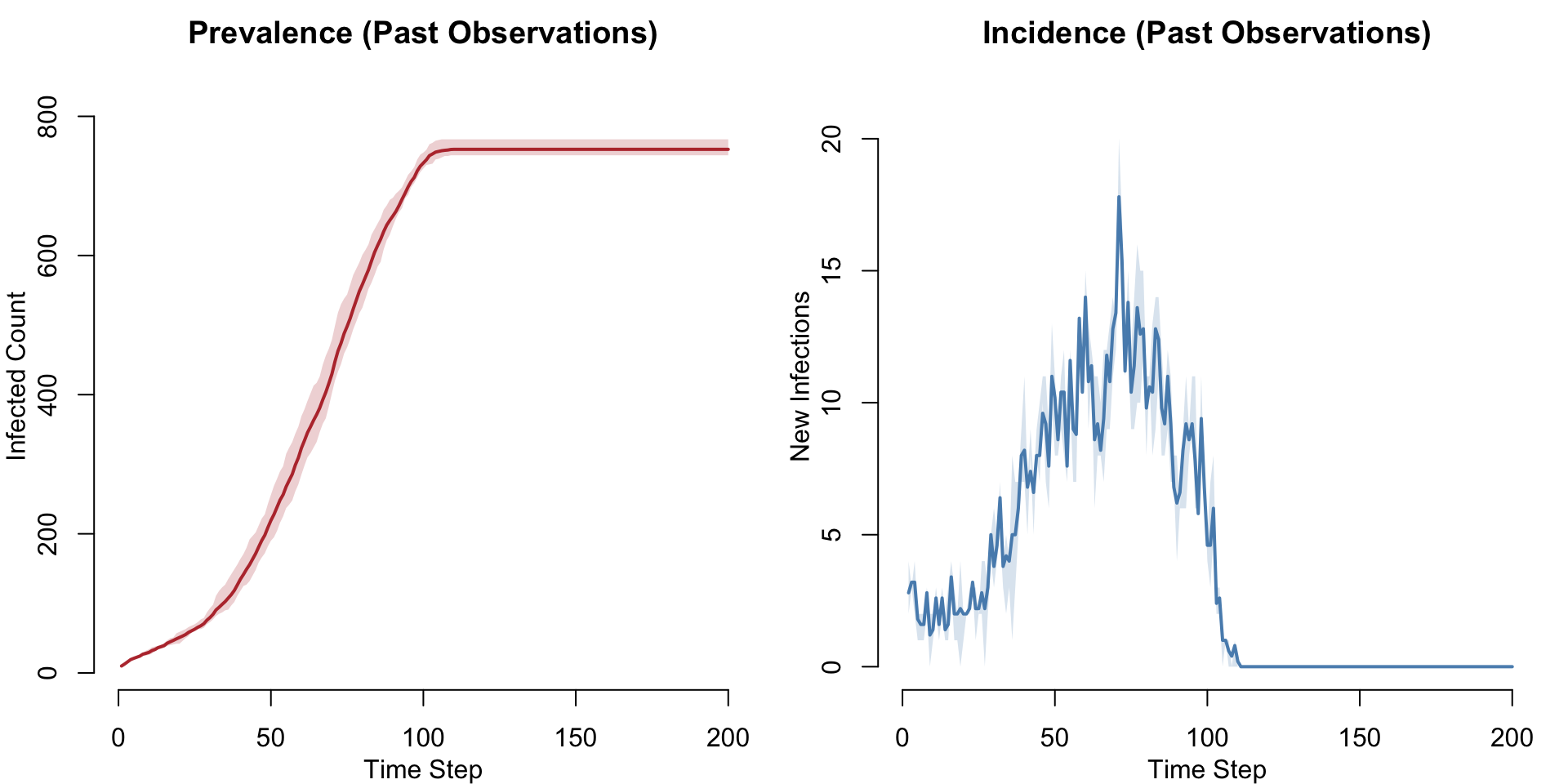
The takeaway: always match nsteps to the number of observed time steps in the network.
Example 2: Time-Varying Transmission
The second scenario models a two-stage disease where transmission probability depends on infection duration. During the primary stage (first 5 timesteps after infection), transmission probability is low (5% per act); during the secondary stage, it increases (15% per act). The track.nw.attr = TRUE parameter stores time-varying disease status on the network, enabling network visualization.
param <- param.net(
inf.prob.stage1 = 0.05,
inf.prob.stage2 = 0.15,
dur.stage1 = 5,
act.rate = 1,
track.nw.attr = TRUE
)
init <- init.net(i.num = 10)control <- control.net(
type = NULL,
nsteps = nsteps,
nsims = nsims,
ncores = ncores,
initialize.FUN = init_obsnw,
infection.FUN = infect_obsnw,
prevalence.FUN = prevalence.net,
resimulate.network = FALSE,
save.nwstats = FALSE,
verbose = FALSE
)sim2 <- netsim(nw, param, init, control)
print(sim2)EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type:
No. simulations: 5
No. time steps: 100
No. NW groups: 1
Fixed Parameters
---------------------------
act.rate = 1
inf.prob.stage1 = 0.05
inf.prob.stage2 = 0.15
dur.stage1 = 5
track.nw.attr = TRUE
groups = 1
Model Functions
-----------------------
initialize.FUN
resim_nets.FUN
summary_nets.FUN
infection.FUN
nwupdate.FUN
prevalence.FUN
verbose.FUN
Model Output
-----------------------
Variables: s.num i.num num si.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5Network Visualization
Because track.nw.attr = TRUE, we can visualize the network colored by disease status at different time points.
par(mfrow = c(1, 2), mar = c(1, 1, 1, 1))
plot(sim2, type = "network", col.status = TRUE, at = 2, sims = 1)
plot(sim2, type = "network", col.status = TRUE, at = nsteps, sims = 1)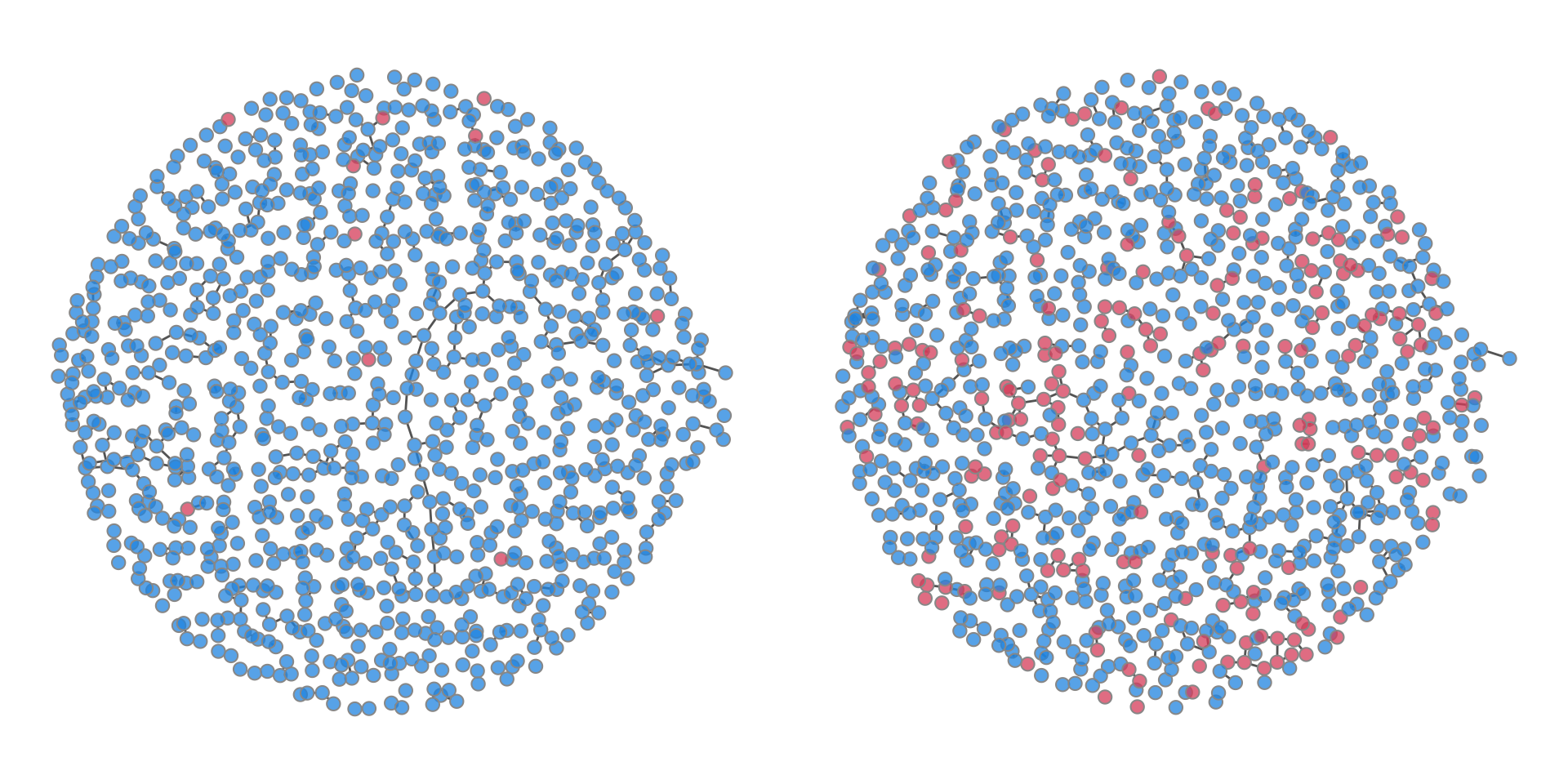
Extracting Individual-Level Data
The time-varying disease status stored on the network can be extracted at any time point using the networkDynamic API:
nwd <- get_network(sim2, 1)
head(get.vertex.attribute.active(nwd, "testatus", at = 1), 25) [1] "s" "s" "s" "s" "s" "s" "s" "s" "s" "s" "s" "s" "s" "s" "s" "s" "s" "s" "s"
[20] "s" "s" "s" "s" "s" "s"head(get.vertex.attribute.active(nwd, "testatus", at = nsteps), 25) [1] "s" "s" "s" "s" "s" "s" "s" "s" "s" "i" "s" "s" "s" "s" "s" "s" "s" "s" "s"
[20] "s" "i" "i" "i" "s" "s"Analysis
Prevalence Comparison
sim <- mutate_epi(sim, prev = i.num / num)
sim2 <- mutate_epi(sim2, prev = i.num / num)par(mfrow = c(1, 2), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim, y = "i.num",
main = "Ex 1: Constant Transmission",
ylab = "Infected Count", xlab = "Time Step",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim2, y = "i.num",
main = "Ex 2: Time-Varying Transmission",
ylab = "Infected Count", xlab = "Time Step",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)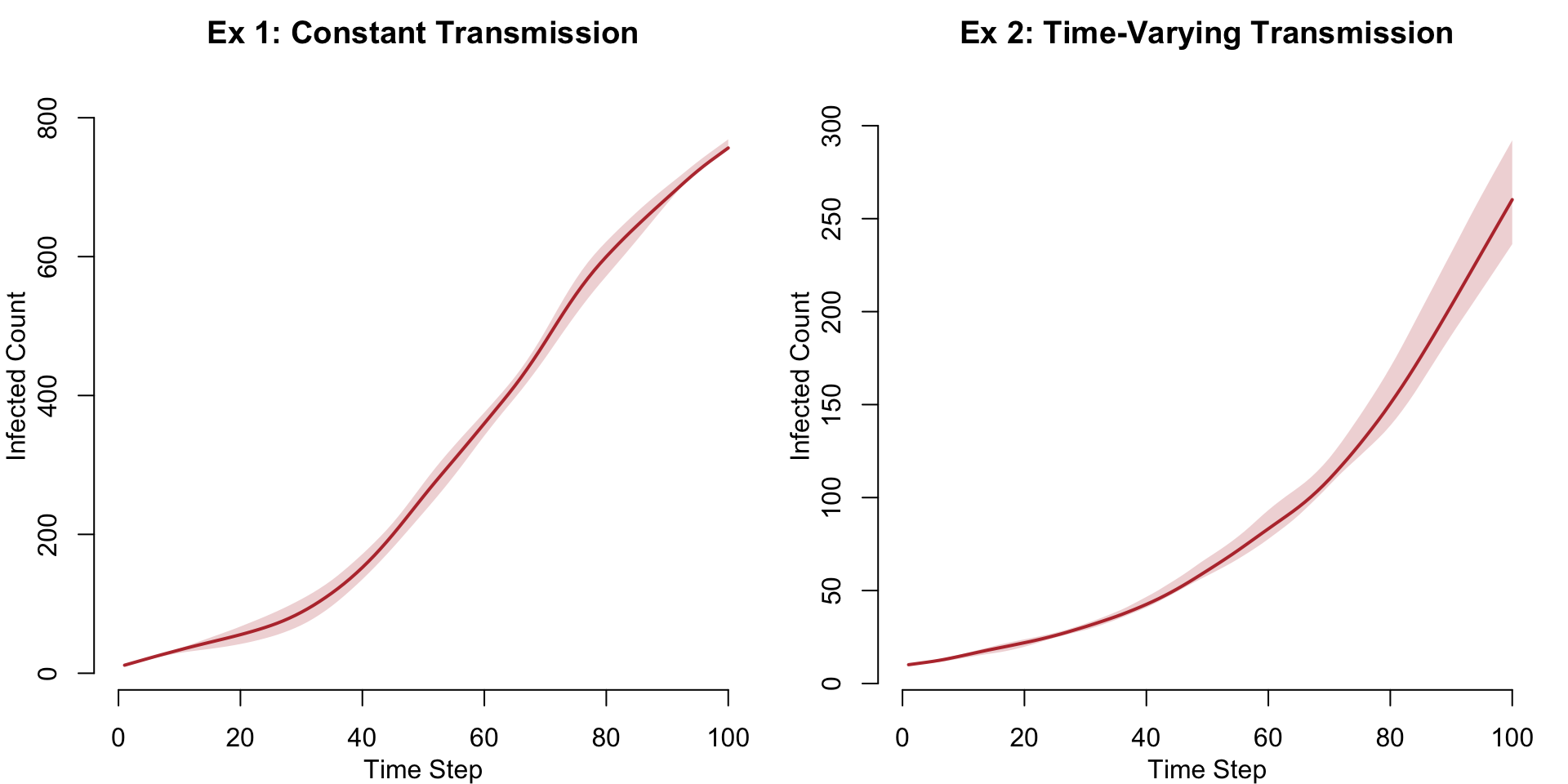
Incidence Comparison
par(mfrow = c(1, 2), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim, y = "si.flow",
main = "Ex 1: Incidence",
ylab = "New Infections", xlab = "Time Step",
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim2, y = "si.flow",
main = "Ex 2: Incidence",
ylab = "New Infections", xlab = "Time Step",
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)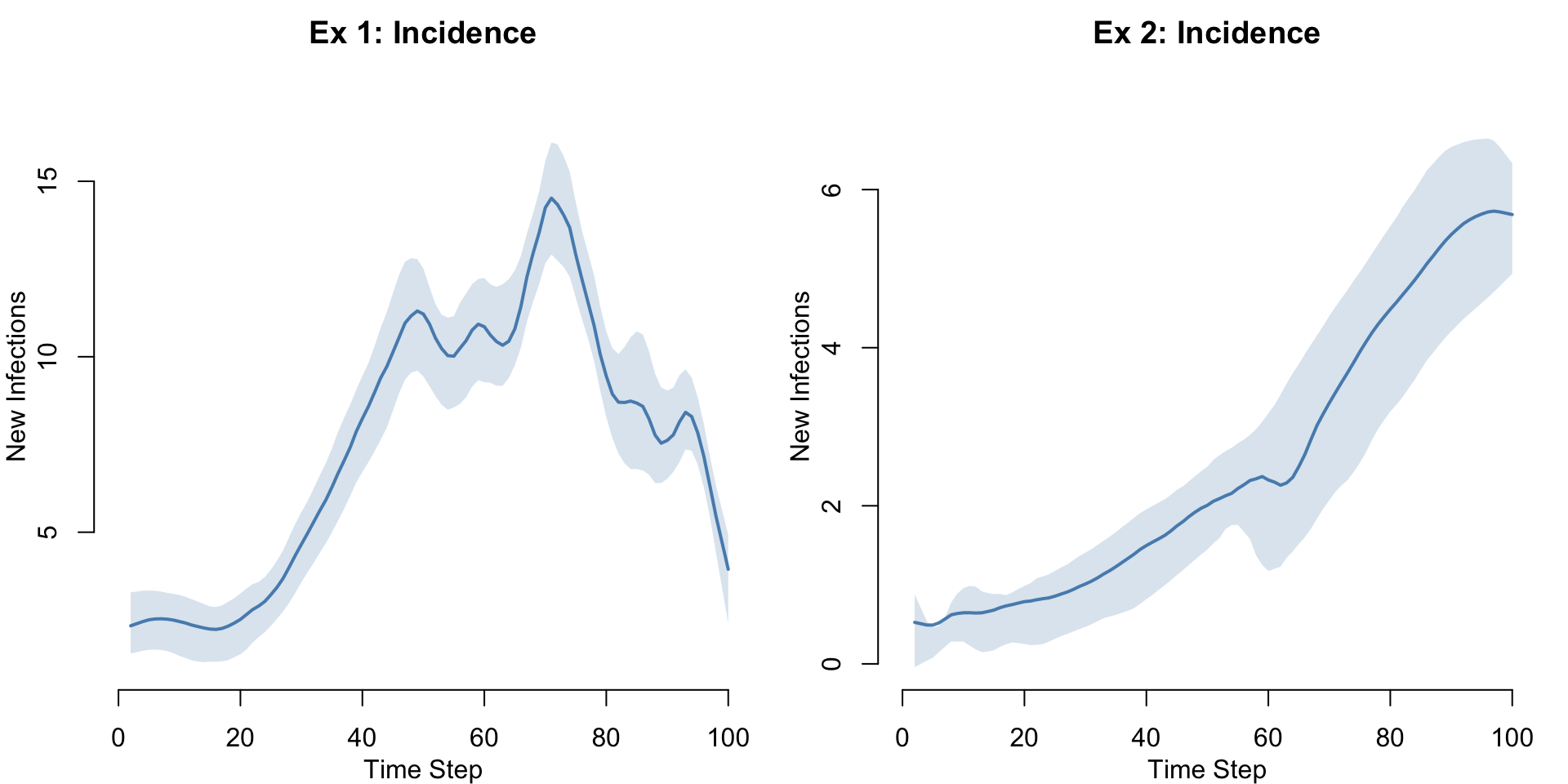
Summary Table
df1 <- as.data.frame(sim)
df2 <- as.data.frame(sim2)
data.frame(
Metric = c("Final prevalence",
"Cumulative infections",
"Mean incidence per step"),
Example1 = c(
round(mean(df1$prev[df1$time == max(df1$time)], na.rm = TRUE), 3),
round(mean(tapply(df1$si.flow, df1$sim, sum, na.rm = TRUE))),
round(mean(df1$si.flow, na.rm = TRUE), 2)
),
Example2 = c(
round(mean(df2$prev[df2$time == max(df2$time)], na.rm = TRUE), 3),
round(mean(tapply(df2$si.flow, df2$sim, sum, na.rm = TRUE))),
round(mean(df2$si.flow, na.rm = TRUE), 2)
)
) Metric Example1 Example2
1 Final prevalence 0.752 0.285
2 Cumulative infections 742.000 275.000
3 Mean incidence per step 7.500 2.780Parameters
Example 1: Basic SI
| Parameter | Description | Value |
|---|---|---|
inf.prob |
Per-act transmission probability | 0.5 |
act.rate |
Acts per partnership per timestep | 1 |
i.num |
Initial number of infected nodes | 10 |
Example 2: Time-Varying Transmission
| Parameter | Description | Value |
|---|---|---|
inf.prob.stage1 |
Transmission probability during primary stage | 0.05 |
inf.prob.stage2 |
Transmission probability during secondary stage | 0.15 |
dur.stage1 |
Duration of primary stage (timesteps) | 5 |
act.rate |
Acts per partnership per timestep | 1 |
track.nw.attr |
Track time-varying disease status on network for visualization | TRUE |
i.num |
Initial number of infected nodes | 10 |
Module Execution Order
At each timestep, modules execute in the following order:
| Order | Module | Function | Description |
|---|---|---|---|
| 1 | Initialization | init_obsnw |
Set up data structure, place observed network, assign initial infections (timestep 1 only) |
| 2 | Infection | infect_obsnw |
Identify discordant partnerships, compute transmission, update infection status |
| 3 | Prevalence | prevalence.net |
Record compartment sizes and flows (built-in EpiModel function) |
Note that unlike most gallery examples, there is no network resimulation step. The resimulate.network = FALSE setting means the observed network is used as-is at every timestep, preserving the exact partnership structure from the census data.
Next Steps
- Add recovery or immunity: Convert this to an SIS or SIR model by adding a recovery module — see the SEIR with Exposed State example for how additional compartments are implemented
- Add vital dynamics: Introduce births and deaths to study longer-term epidemics — see SI with Vital Dynamics
- Use a different dataset: The
networkDynamicDatapackage contains several other observed dynamic networks that could be substituted - Bridge approaches: Take a static network dataset (e.g., from the
ergmpackage) and fit a TERGM with an assumed edge dissolution rate, combining observed and model-based methods