flowchart LR
S["<b>S</b><br/>Susceptible"] -->|"infection<br/>(si.flow)"| I["<b>I</b><br/>Infectious"]
style S fill:#3498db,color:#fff
style I fill:#e74c3c,color:#fff
Epidemics with Multiple (Multilayer) Networks
Overview
This example demonstrates how to simulate an epidemic on a multilayer network in EpiModel. A multilayer network consists of two or more network layers that share the same node set but have different edge sets — representing distinct types of relationships. For example, the two layers might represent main partnerships and casual contacts, or sexual contacts and needle-sharing contacts.
The key feature of multilayer models is cross-layer dependency: the degree (number of edges) in one layer can influence edge formation in the other. This models the realistic constraint that individuals have finite relational capacity — being active in one type of partnership reduces availability for the other. The cross-layer dependency is maintained during simulation by a callback function that updates degree attributes between layer resimulations.
This example uses a simple SI model in a closed population to isolate the multilayer network mechanics from disease model complexity. No custom module-fx.R is needed — it uses EpiModel’s built-in SI modules. The multilayer functionality is entirely in the network estimation and simulation layers.
- model.R — Main simulation script (no module-fx.R needed for this example)
Model Structure
| Compartment | Label | Description |
|---|---|---|
| Susceptible | S | Not infected; at risk of infection through contact on either network layer |
| Infectious | I | Infected and capable of transmitting (no recovery in SI) |
Network Layers
The two network layers have different formation models, target statistics, and mean durations. Transmission can occur over edges in either layer.
| Property | Layer 1 | Layer 2 |
|---|---|---|
| Interpretation | Longer-duration partnerships | Shorter-duration partnerships |
| Target edges | 90 | 75 |
| Mean degree | 0.36 | 0.30 |
| Mean duration | 100 time steps | 75 time steps |
| Formation terms | edges + nodematch("race") + nodefactor("deg1+.net2") |
edges + degree(1) + nodefactor("deg1+.net1") |
| Cross-layer term | nodefactor("deg1+.net2") |
nodefactor("deg1+.net1") |
Cross-Layer Dependency
The nodefactor("deg1+.netX") terms create negative degree correlation across layers. The target statistic for these terms (10 out of 90 or 75 total edges) is deliberately small relative to total edges, producing strong negative correlation: most nodes that are active in one layer are inactive in the other.
Setup
suppressMessages(library(EpiModel))
nsims <- 5
ncores <- 5
nsteps <- 500Network Model
Network Initialization
Initialize a network of 500 nodes with a binary demographic attribute (race). This attribute is used for assortative mixing (homophily) in layer 1’s formation model.
n <- 500
nw <- network_initialize(n)
nw <- set_vertex_attribute(nw, "race", rep(0:1, length.out = n))Generating Starting Degree Values with san()
Because the two layers depend on each other’s degree distribution (via cross-layer nodefactor terms), we need reasonable starting values for the degree attributes before fitting the ERGM models. We use san() (simulated annealing from the ergm package) to generate edge sets consistent with our target statistics.
Step 1: Generate layer 1 edges consistent with target mean degree and race homophily. Layer 1 has target stats of 90 total edges and 60 that are race-homophilous (same-race partnerships).
- 1
- Record each node’s exact degree in layer 1 as a vertex attribute.
- 2
- Create a binary indicator (0 vs. 1+) for the cross-layer formation term.
Step 2: Generate layer 2 edges, accounting for the cross-layer constraint from step 1. The nodefactor("deg1+.net1") term ensures that nodes already active in layer 1 are less likely to form layer 2 edges (negative correlation).
nw_san_2 <- san(nw_san ~ edges + degree(1) + nodefactor("deg1+.net1")
+ Sum(matrix(seq_len(network.size(nw)), nrow = 1) ~
degrange(1, by = "deg.net1", levels = -1),
label = "Sum"),
1 target.stats = c(75, 120, 10, 10))
nw_san <- set_vertex_attribute(nw_san, "deg1+.net2",
pmin(get_degree(nw_san_2), 1))- 1
-
Targets: 75 total edges, 120 degree-1 nodes, 10 edges involving nodes active in layer 1, and 10 for the
Sumterm linking layer 2 degree to layer 1 degree categories.
Verify that both SAN networks approximate their target statistics. These do not need to be exact — they are starting conditions for the ERGM estimation.
# Layer 1 targets: 90 edges, 60 homophilous
summary(nw_san ~ edges + nodematch("race") + nodefactor("deg1+.net2")) edges nodematch.race nodefactor.deg1+.net2.1
90 60 10 # Layer 2 targets: 75 edges, 120 degree-1
summary(nw_san_2 ~ edges + degree(1) + nodefactor("deg1+.net1")) edges degree1 nodefactor.deg1+.net1.1
75 120 10 Set the binary degree attributes on the original network object for estimation.
nw <- set_vertex_attribute(nw, "deg1+.net1", pmin(get_degree(nw_san), 1))
nw <- set_vertex_attribute(nw, "deg1+.net2", pmin(get_degree(nw_san_2), 1))Network Model Estimation
The formation models include both layer-specific terms and cross-layer dependency terms. Layer 1 models race homophily; layer 2 models the degree-1 count. Both include a nodefactor term for the other layer’s degree, creating negative degree correlation across layers.
- 1
- Layer 1: 90 edges total, 60 race-homophilous, 10 among nodes with layer-2 edges (strong negative correlation).
- 2
- Layer 2: 75 edges total, 120 degree-1 nodes, 10 among nodes with layer-1 edges.
Dissolution models specify the mean partnership duration for each layer independently.
- 1
- Layer 1: longer, more stable partnerships (mean duration 100 time steps).
- 2
- Layer 2: shorter, more transient partnerships (mean duration 75 time steps).
Although we think of this as one multilayer network, estimation is done separately for each layer. The layers are linked through the shared nodal attributes (deg1+.net1, deg1+.net2).
est.1 <- netest(nw, formation.1, target.stats.1, coef.diss.1)
est.2 <- netest(nw, formation.2, target.stats.2, coef.diss.2)Network Diagnostics
Run diagnostics for each layer. Because the network is relatively small (500 nodes) and some target statistics are small (10 for cross-layer terms), there may be more variability than in a single-layer model.
dx.1 <- netdx(est.1, nsims = nsims, ncores = ncores, nsteps = nsteps)
Network Diagnostics
-----------------------
- Simulating 5 networks
- Calculating formation statisticsprint(dx.1)EpiModel Network Diagnostics
=======================
Diagnostic Method: Dynamic
Simulations: 5
Time Steps per Sim: 500
Formation Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means)
edges 90 88.612 -1.542 1.032 -1.344 3.101
nodematch.race 60 60.950 1.583 0.991 0.958 4.306
nodefactor.deg1+.net2.1 10 10.928 9.280 0.603 1.540 2.323
SD(Statistic)
edges 6.793
nodematch.race 6.476
nodefactor.deg1+.net2.1 3.362
Duration Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 100 98.604 -1.396 1.742 -0.801 5.678 9.827
Dissolution Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.01 0.01 -3.571 0 -1.734 0 0.01plot(dx.1)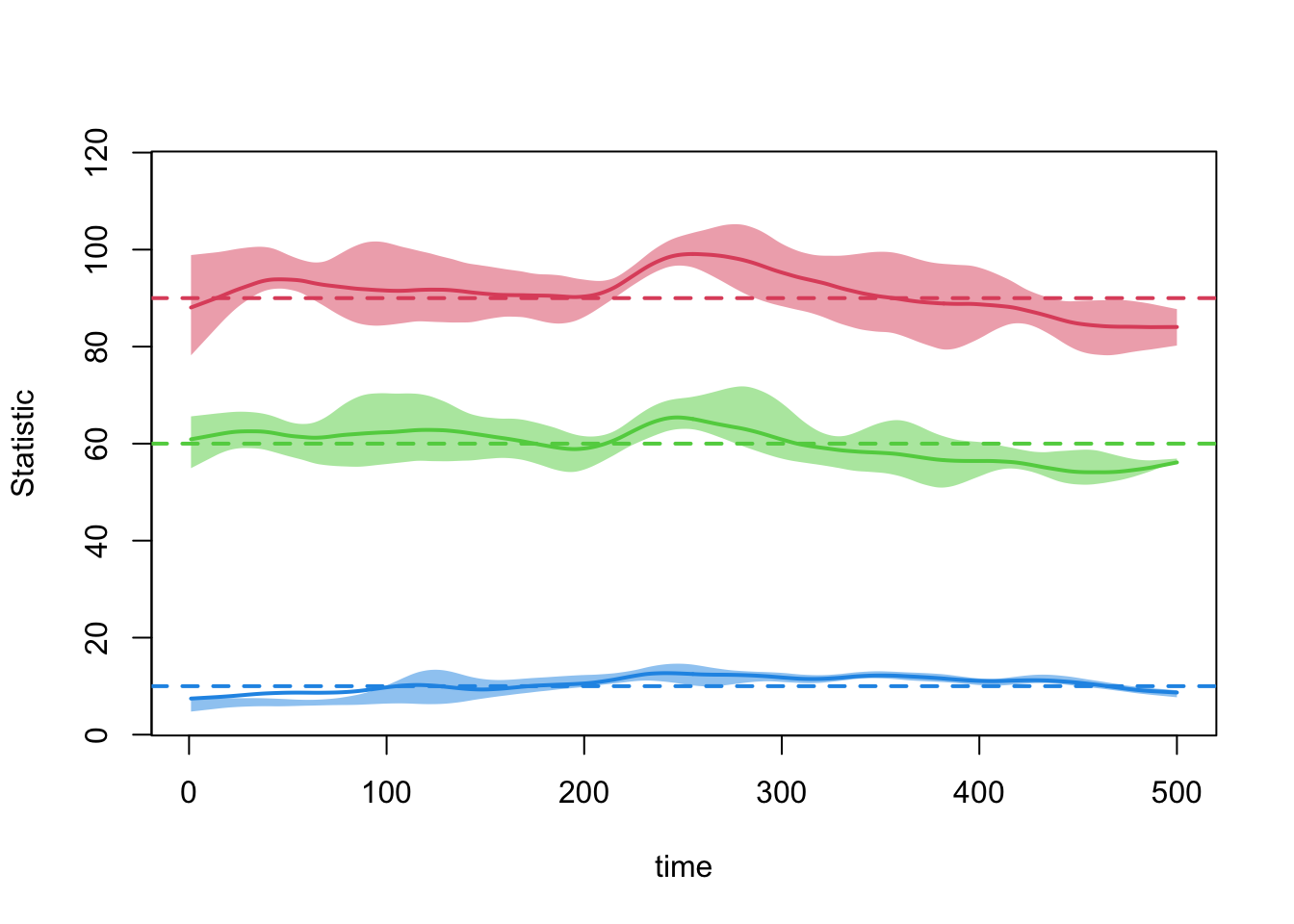
dx.2 <- netdx(est.2, nsims = nsims, ncores = ncores, nsteps = nsteps)
Network Diagnostics
-----------------------
- Simulating 5 networks
- Calculating formation statisticsprint(dx.2)EpiModel Network Diagnostics
=======================
Diagnostic Method: Dynamic
Simulations: 5
Time Steps per Sim: 500
Formation Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means)
edges 75 76.048 1.398 1.108 0.946 1.788
degree1 120 118.266 -1.445 1.547 -1.121 2.777
nodefactor.deg1+.net1.1 10 11.005 10.052 0.622 1.616 1.685
SD(Statistic)
edges 6.953
degree1 10.080
nodefactor.deg1+.net1.1 3.218
Duration Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 75 78.181 4.242 1.535 2.073 3.463 8.179
Dissolution Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.013 0.013 -0.54 0 -0.256 0 0.013plot(dx.2)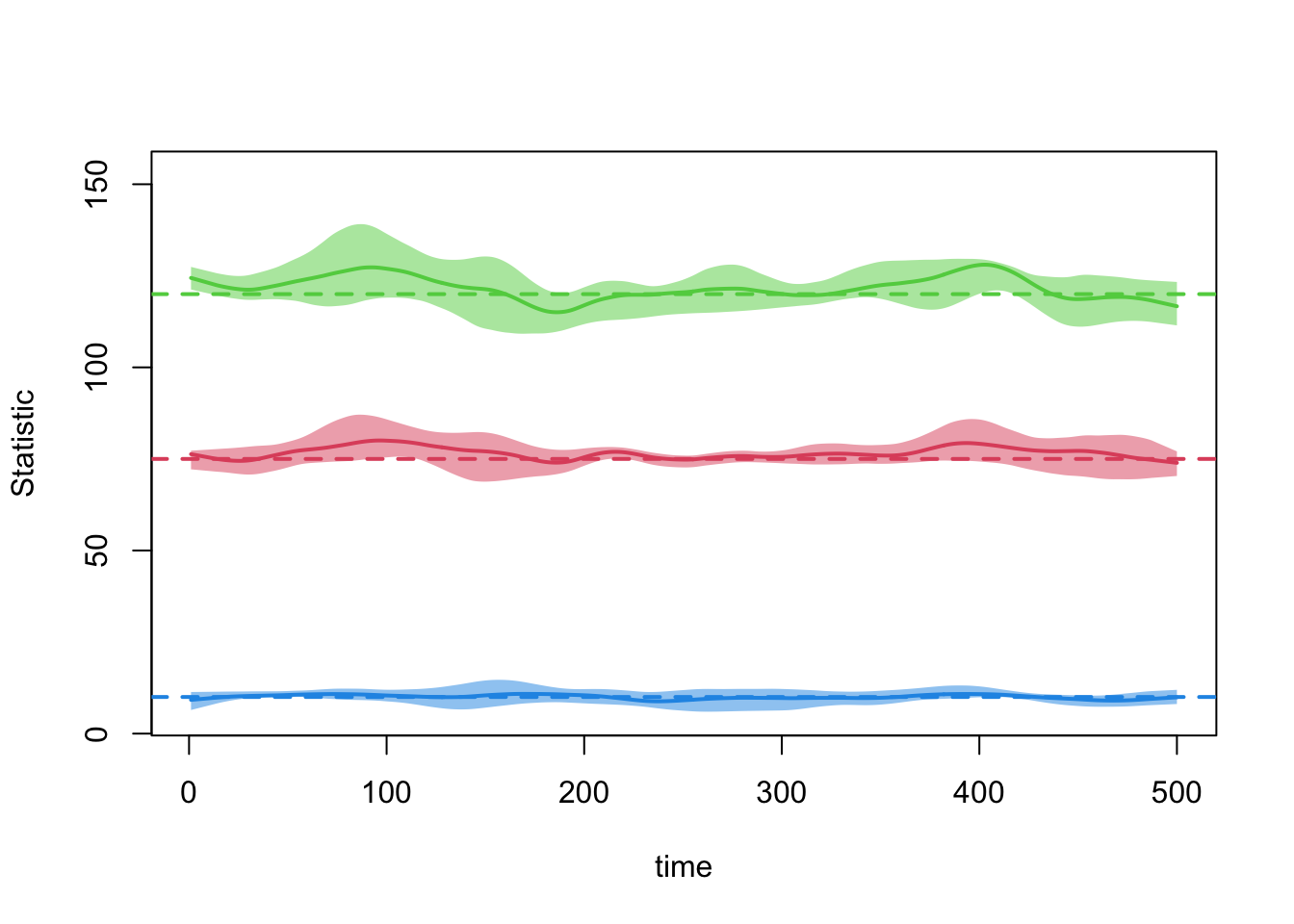
Epidemic Simulation
Parameters
Simple SI model parameters. The high transmission probability and act rate are deliberate — they generate a clearly visible epidemic so we can focus on the network mechanics rather than epidemiological realism.
Cross-Layer Update Function
This is the key mechanism for multilayer feedback. During each time step, EpiModel resimulates each network layer sequentially. Before resimulating layer k, it calls this function to update the cross-layer degree attributes that layer k’s formation model depends on. Without this function, the cross-layer degree attributes would be frozen at their initial values and the layers would evolve independently.
network_layer_updates <- function(dat, at, network) {
if (network == 0) {
1 dat <- set_attr(dat, "deg1+.net2", pmin(get_degree(dat$run$el[[2]]), 1))
} else if (network == 1) {
2 dat <- set_attr(dat, "deg1+.net1", pmin(get_degree(dat$run$el[[1]]), 1))
} else if (network == 2) {
3 dat <- set_attr(dat, "deg1+.net2", pmin(get_degree(dat$run$el[[2]]), 1))
}
return(dat)
}- 1
-
network == 0(before layer 1 resimulation): updatedeg1+.net2from current layer 2 edges. - 2
-
network == 1(before layer 2 resimulation): updatedeg1+.net1from current layer 1 edges. - 3
-
network == 2(after layer 2 resimulation): recalculatedeg1+.net2for summary statistics.
Control Settings and Simulation
control <- control.net(
type = "SI",
nsteps = nsteps,
nsims = nsims,
ncores = ncores,
1 tergmLite = TRUE,
2 resimulate.network = TRUE,
3 dat.updates = network_layer_updates,
4 nwstats.formula = multilayer(
"formation",
~edges + nodefactor("deg1+.net1") + degree(0:4)
)
)
5sim <- netsim(list(est.1, est.2), param, init, control)- 1
-
Use lightweight edgelist representation (required for
dat$run$el[[]]access in the update callback). - 2
- Redraw both layers each time step.
- 3
- The cross-layer degree update callback runs between layer resimulations.
- 4
-
multilayer()specifies per-layer network statistics to track. Layer 1 uses the default formation formula; layer 2 additionally tracks the full degree distribution (degree 0 through 4). - 5
-
Passing a list of
netestobjects tells EpiModel this is a multilayer model. The list order determines layer numbering.
Analysis
Network Formation During Simulation
Verify that network structure is maintained during the epidemic. These plots should show the simulated formation statistics tracking close to their targets throughout the run.
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
print(sim, network = 1)EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type: SI
No. simulations: 5
No. time steps: 500
No. NW groups: 1
Fixed Parameters
---------------------------
inf.prob = 0.5
act.rate = 2
groups = 1
Model Output
-----------------------
Variables: s.num i.num num si.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means)
edges 90 95.111 5.679 2.446 2.090 7.621
nodematch.race 60 63.020 5.033 1.757 1.719 2.938
nodefactor.deg1+.net2.1 10 12.030 20.300 0.799 2.541 2.211
SD(Statistic)
edges 11.476
nodematch.race 8.344
nodefactor.deg1+.net2.1 4.510
Duration and Dissolution Statistics
-----------------------
Not available when:
- `control$tergmLite == TRUE`
- `control$save.network == FALSE`
- `control$save.diss.stats == FALSE`
- dissolution formula is not `~ offset(edges)`
- `keep.diss.stats == FALSE` (if merging)plot(sim, network = 1, type = "formation",
main = "Layer 1: Formation Diagnostics During Simulation")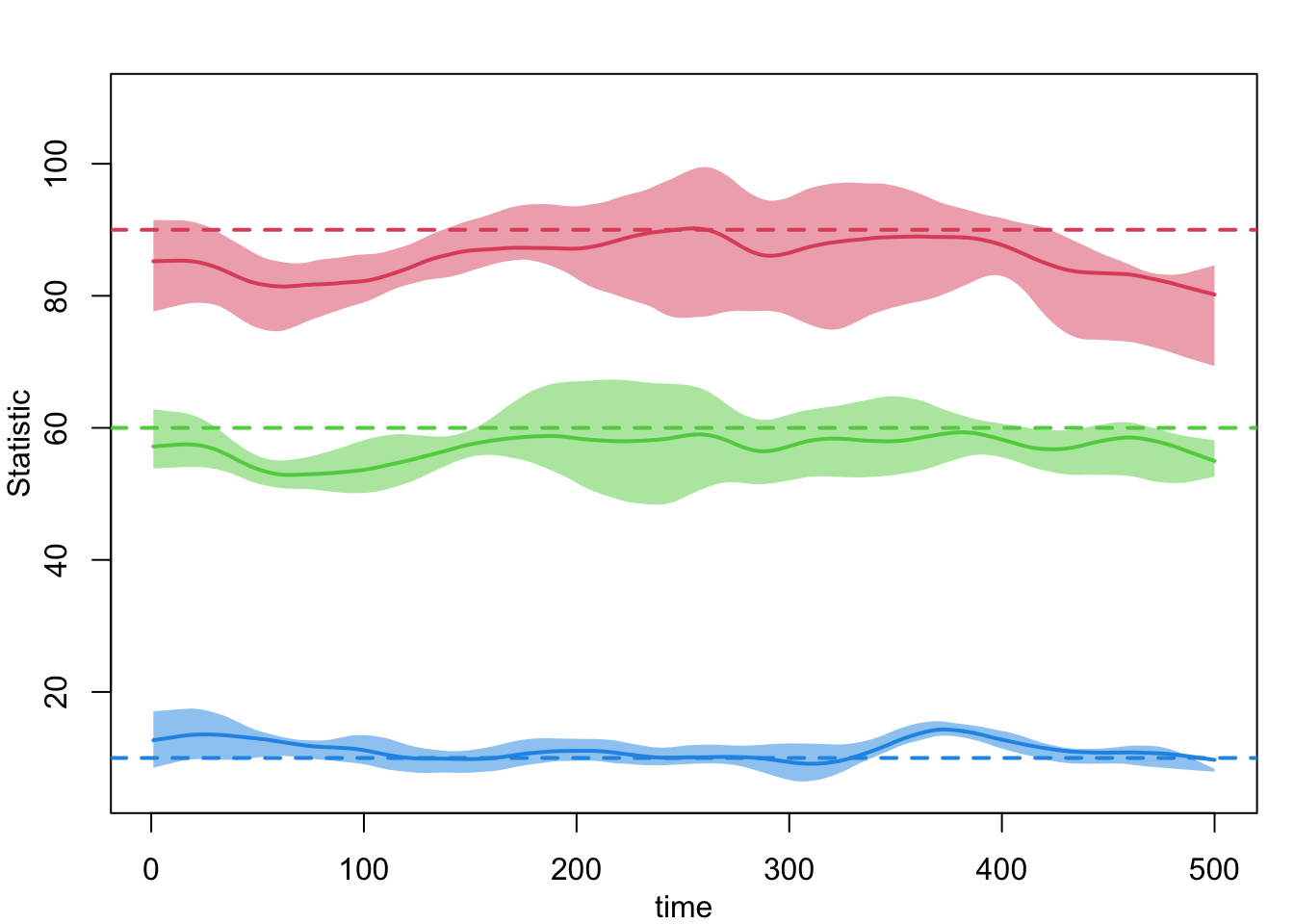
print(sim, network = 2)EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type: SI
No. simulations: 5
No. time steps: 500
No. NW groups: 1
Fixed Parameters
---------------------------
inf.prob = 0.5
act.rate = 2
groups = 1
Model Output
-----------------------
Variables: s.num i.num num si.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means)
edges 75 69.847 -6.871 1.204 -4.280 3.461
nodefactor.deg1+.net1.1 10 11.085 10.848 0.832 1.304 1.764
degree0 NA 374.315 NA 1.914 NA 6.030
degree1 120 112.976 -5.853 1.614 -4.353 5.333
degree2 NA 11.480 NA 0.457 NA 0.745
degree3 NA 1.166 NA 0.142 NA 0.154
degree4 NA 0.055 NA 0.010 NA 0.063
SD(Statistic)
edges 7.129
nodefactor.deg1+.net1.1 4.390
degree0 11.923
degree1 10.693
degree2 3.285
degree3 1.038
degree4 0.229
Duration and Dissolution Statistics
-----------------------
Not available when:
- `control$tergmLite == TRUE`
- `control$save.network == FALSE`
- `control$save.diss.stats == FALSE`
- dissolution formula is not `~ offset(edges)`
- `keep.diss.stats == FALSE` (if merging)plot(sim, network = 2, type = "formation",
main = "Layer 2: Formation Diagnostics During Simulation")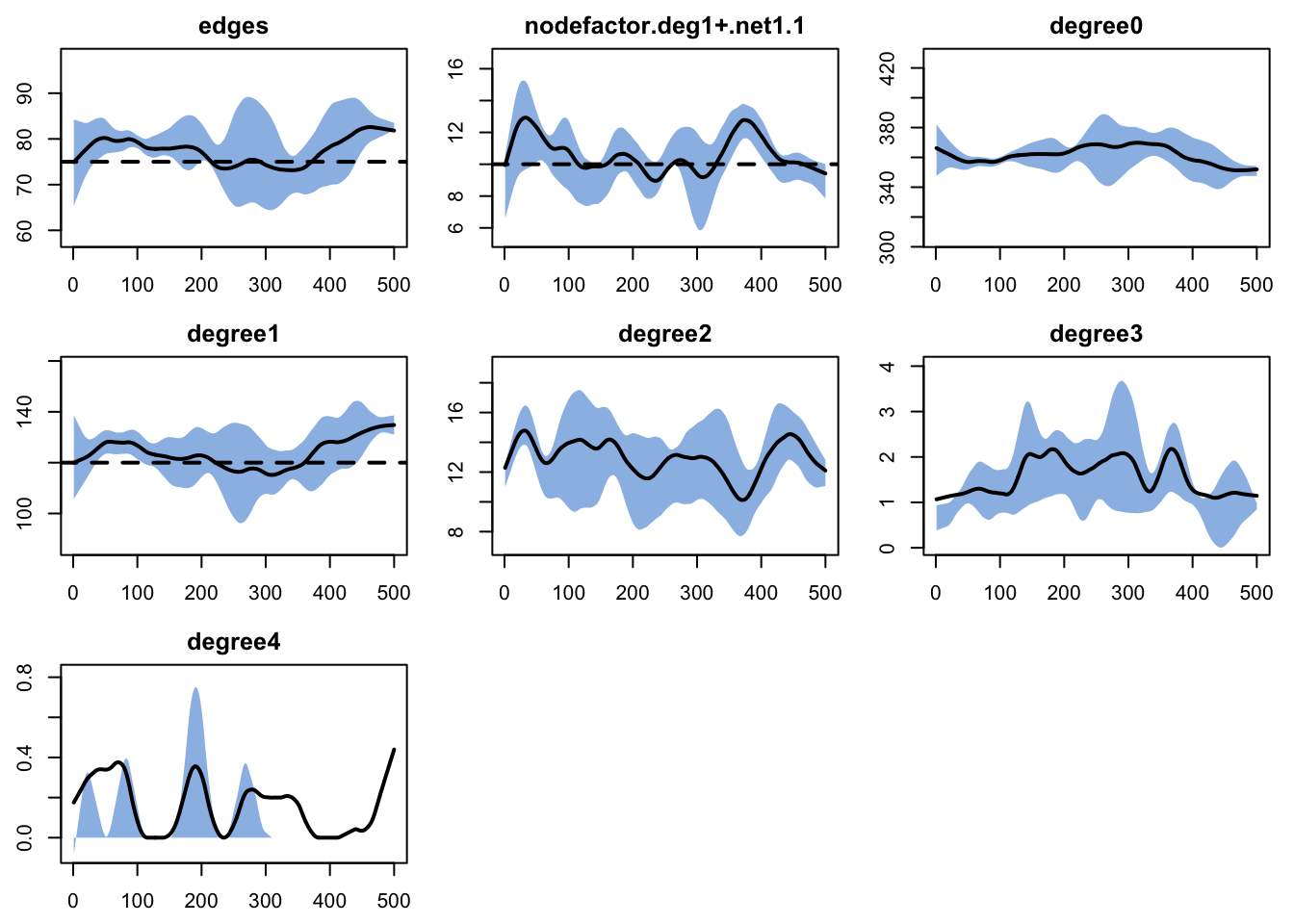
SI Compartment Counts
plot(sim, y = c("s.num", "i.num"),
main = "SI Epidemic on Multilayer Network",
ylab = "Count", xlab = "Time Steps",
mean.col = c("steelblue", "firebrick"),
mean.lwd = 2, mean.smooth = TRUE,
qnts.col = c("steelblue", "firebrick"),
qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = TRUE)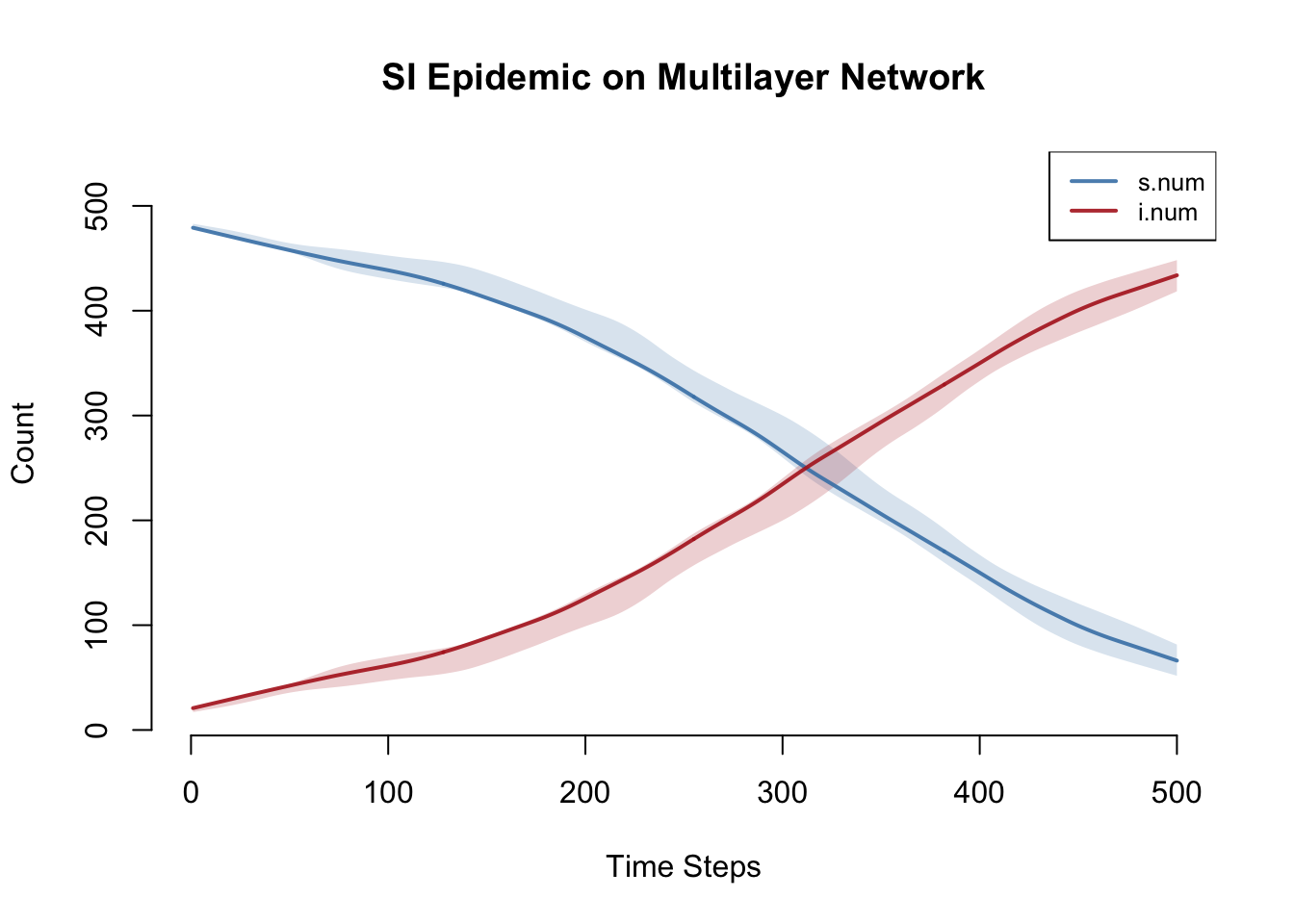
Prevalence
sim <- mutate_epi(sim, prev = i.num / num)
plot(sim, y = "prev",
main = "Prevalence over Time",
ylab = "Prevalence", xlab = "Time Steps",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE)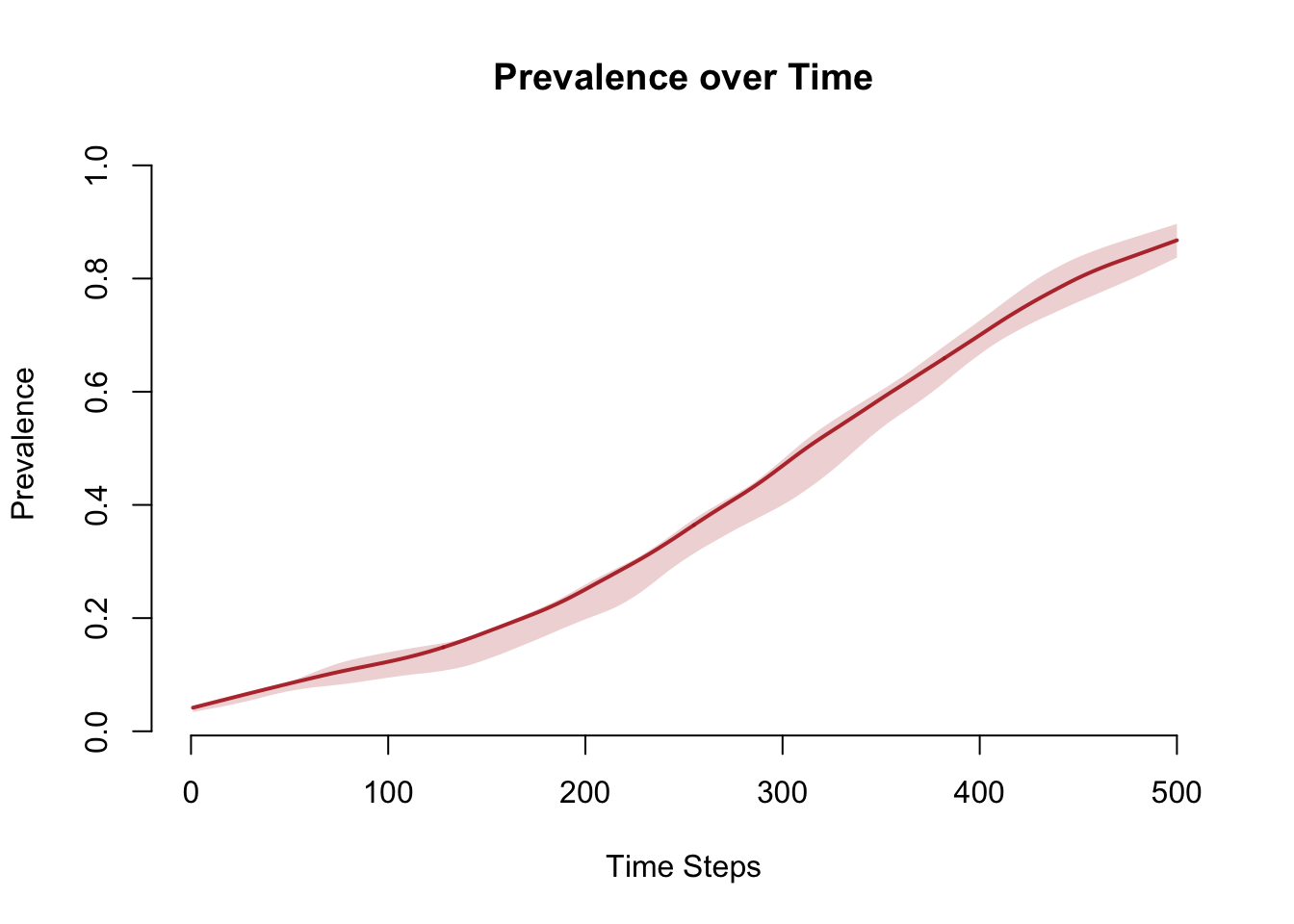
Incidence
plot(sim, y = "si.flow",
main = "New Infections per Time Step",
ylab = "Incidence", xlab = "Time Steps",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE)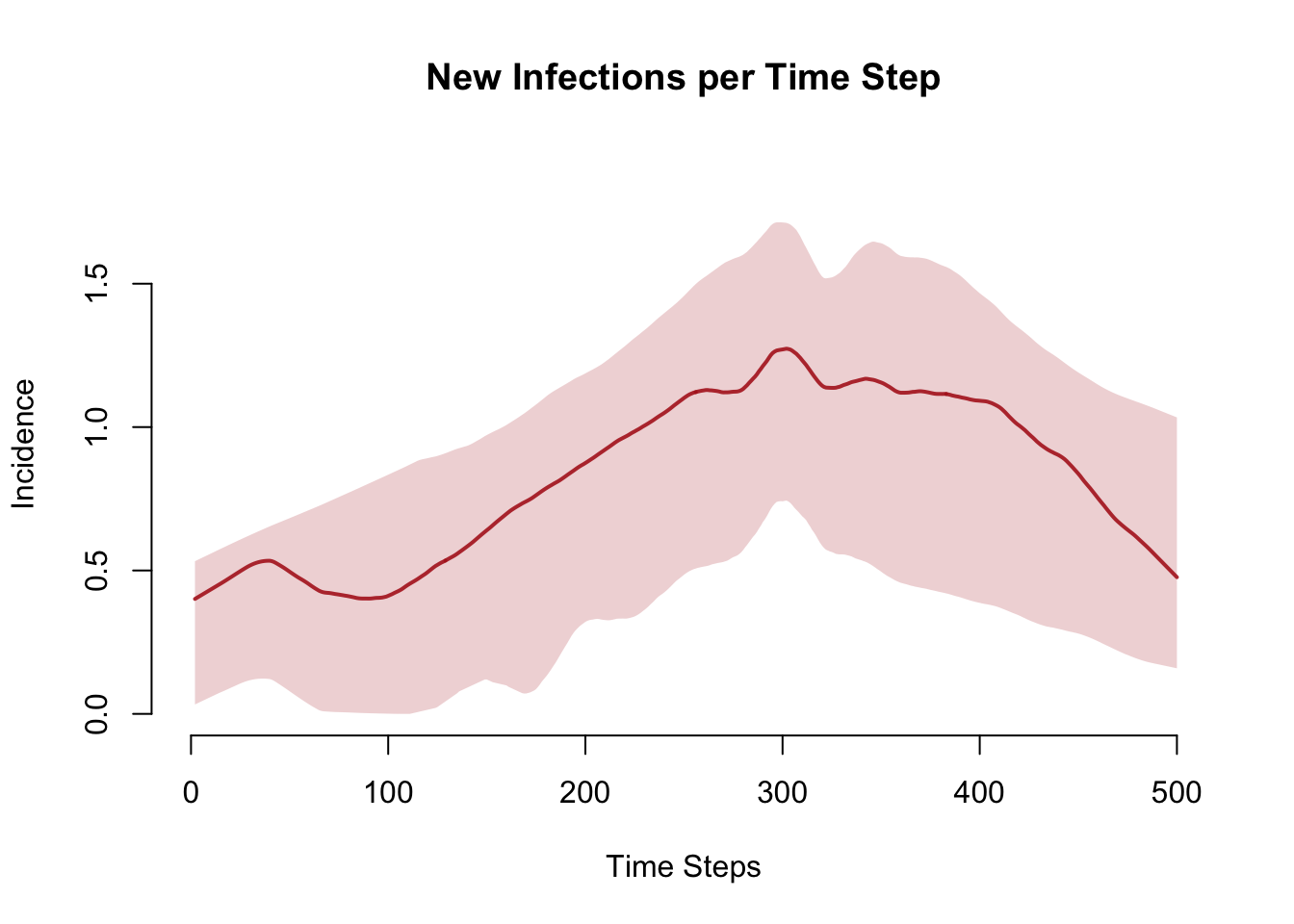
Summary Table
df <- as.data.frame(sim)
last_t <- max(df$time)
knitr::kable(data.frame(
Metric = c("Population size",
"Final susceptible",
"Final infected",
"Final prevalence",
"Cumulative new infections",
"Peak incidence (per time step)"),
Value = c(n,
round(mean(df$s.num[df$time == last_t])),
round(mean(df$i.num[df$time == last_t])),
round(mean(df$prev[df$time == last_t]), 3),
round(mean(tapply(df$si.flow, df$sim, sum, na.rm = TRUE))),
max(df$si.flow, na.rm = TRUE))
))| Metric | Value |
|---|---|
| Population size | 500.000 |
| Final susceptible | 63.000 |
| Final infected | 437.000 |
| Final prevalence | 0.874 |
| Cumulative new infections | 427.000 |
| Peak incidence (per time step) | 9.000 |
Parameters
Epidemic
| Parameter | Description | Value |
|---|---|---|
inf.prob |
Per-act transmission probability | 0.5 |
act.rate |
Acts per partnership per time step | 2 |
Network Formation
| Parameter | Layer 1 | Layer 2 |
|---|---|---|
| Target edges | 90 | 75 |
Race homophily (nodematch) |
60 | — |
Degree-1 count (degree(1)) |
— | 120 |
Cross-layer nodefactor |
10 | 10 |
Network Dissolution
| Parameter | Layer 1 | Layer 2 |
|---|---|---|
| Mean duration (time steps) | 100 | 75 |
Key EpiModel Functions for Multilayer Models
| Function / Argument | Purpose |
|---|---|
netsim(list(est.1, est.2), ...) |
Passing a list of netest objects signals a multilayer model |
dat.updates in control.net() |
Callback function for cross-layer attribute updates |
multilayer() in nwstats.formula |
Specifies per-layer network statistics to track |
tergmLite = TRUE |
Uses lightweight edgelist representation (required for dat$run$el[[]] access) |
resimulate.network = TRUE |
Required for multilayer models with cross-layer dependency |
Next Steps
- Layer-specific transmission: different
inf.proboract.rateby edge type (e.g., lower transmission in casual vs. main partnerships) - More than two layers: add a third network layer (e.g., needle-sharing)
- Vital dynamics: add arrivals and departures with custom modules — see SI with Vital Dynamics for the departure and arrival module pattern
- Different disease models: extend to SIR, SEIR, or disease-specific models — see Adding an Exposed State for the progression module pattern
- Heterogeneous populations: add more nodal attributes beyond the binary
raceattribute used here