flowchart TD
S["<b>S</b><br/>Susceptible"] -->|"infection<br/>(si.flow)"| I_undx
subgraph Infected ["Infectious (I)"]
I_undx["Undiagnosed<br/>rec.rate = 0.05"] -->|"testing<br/>(test.rate)"| I_dx["Diagnosed<br/>rec.rate.tx = 0.5"]
I_dx -->|"test.dur expires"| I_undx
end
I_undx -->|"natural clearance<br/>(slow)"| S
I_dx -->|"treatment<br/>(fast)"| S
style S fill:#3498db,color:#fff
style I_undx fill:#e74c3c,color:#fff
style I_dx fill:#f39c12,color:#fff
Test and Treat Intervention for an SIS Epidemic
Overview
This example demonstrates how to build a test-and-treat intervention for a bacterial STI (e.g., gonorrhea) modeled as an SIS epidemic over a dynamic network. Many STIs are asymptomatic — infected individuals do not know they are infected without routine screening. Testing identifies infected individuals who then receive antibiotic treatment, dramatically accelerating recovery. The key question: how much screening is needed to meaningfully reduce population-level prevalence?
Unlike SIR models where recovered individuals gain immunity, the SIS framework allows reinfection — recovered individuals return to the susceptible pool and can be infected again. This creates an endemic equilibrium where new infections balance recoveries. Interventions that increase the recovery rate (like test-and-treat) shift this equilibrium to a lower prevalence.
This is the foundational intervention example in the Gallery. The techniques shown here — adding custom attributes for diagnosis status, implementing heterogeneous rates based on intervention status, and comparing intervention scenarios — are building blocks for more complex treatment cascade models.
- model.R — Main simulation script
- module-fx.R — Custom module functions
Model Structure
| Compartment | Label | Description |
|---|---|---|
| Susceptible | S | Not infected; at risk of infection (including previously recovered) |
| Infectious | I | Infected; can transmit to susceptible contacts |
Within the infectious compartment, individuals can be either undiagnosed (recovering slowly via natural clearance) or diagnosed (recovering quickly via antibiotic treatment).
In an SIS model, recovered individuals immediately return to the susceptible pool. There is no immune compartment, so the epidemic can persist indefinitely as an endemic equilibrium. Reinfection is common, and interventions must be sustained — unlike SIR vaccination, there is no herd immunity threshold.
The test-and-treat intervention follows a three-step cascade and is implemented with two separate attributes so that “tested” and “diagnosed positive” are not conflated:
- Testing: Eligible individuals (active, not currently in a test window, not on active treatment) test at rate
test.rateper timestep. Testing is universal — both susceptibles and infecteds can test, and it does not require symptoms. Every tester getstested.status = 1fortest.durtimesteps, which gates re-testing. - Positive diagnosis: Only infected testers receive
diag.status = 1. Susceptible testers got a negative result — they havetested.status = 1butdiag.status = 0, and they do not enter the treatment pipeline. - Treatment course: Once positively diagnosed, the individual recovers at rate
rec.rate.txfor up totest.durtimesteps. If they have not recovered by then, the treatment course ends and they revert to the natural clearance rate until re-tested.
On recovery, diag.status is cleared (the individual is cured and exits the treatment pipeline).
Setup
suppressMessages(library(EpiModel))
nsims <- 5
ncores <- 5
nsteps <- 500Custom Modules
Test and Treat Module
This module implements the screening and diagnosis cascade. Two attributes are tracked separately:
tested.statusflags individuals who tested recently (within the lasttest.durtimesteps), regardless of result. It gates re-testing eligibility so the same person isn’t tested every step.diag.statusflags individuals who tested positive and are infected. It gates the treatment-rate recovery in the recovery module.
Universal screening: both susceptibles and infecteds can test. Susceptibles receive a negative result and never enter the treatment pipeline; only infected testers get diag.status = 1.
tnt <- function(dat, at) {
## Attributes ##
active <- get_attr(dat, "active")
status <- get_attr(dat, "status")
1 if (at == 2) {
dat <- set_attr(dat, "tested.status", rep(0, length(active)))
dat <- set_attr(dat, "tested.time", rep(NA, length(active)))
dat <- set_attr(dat, "diag.status", rep(0, length(active)))
dat <- set_attr(dat, "diag.time", rep(NA, length(active)))
}
tested.status <- get_attr(dat, "tested.status")
tested.time <- get_attr(dat, "tested.time")
diag.status <- get_attr(dat, "diag.status")
diag.time <- get_attr(dat, "diag.time")
## Parameters ##
test.rate <- get_param(dat, "test.rate")
test.dur <- get_param(dat, "test.dur")
## Testing process ##
2 idsElig <- which(active == 1 & tested.status == 0 & diag.status == 0)
nElig <- length(idsElig)
3 vecTest <- which(rbinom(nElig, 1, test.rate) == 1)
idsTest <- idsElig[vecTest]
nTest <- length(idsTest)
# Every tester enters the test-window (gates re-testing).
tested.status[idsTest] <- 1
tested.time[idsTest] <- at
4 # Only infected testers receive a positive diagnosis.
idsTestPos <- idsTest[status[idsTest] == "i"]
diag.status[idsTestPos] <- 1
diag.time[idsTestPos] <- at
nDiagNew <- length(idsTestPos)
## Test-window reset ##
5 idsTestedReset <- which(at - tested.time > (test.dur - 1))
tested.status[idsTestedReset] <- 0
tested.time[idsTestedReset] <- NA
## Treatment-course reset ##
6 idsDiagReset <- which(at - diag.time > (test.dur - 1))
diag.status[idsDiagReset] <- 0
diag.time[idsDiagReset] <- NA
## Write out updated attributes ##
dat <- set_attr(dat, "tested.status", tested.status)
dat <- set_attr(dat, "tested.time", tested.time)
dat <- set_attr(dat, "diag.status", diag.status)
dat <- set_attr(dat, "diag.time", diag.time)
## Summary statistics ##
dat <- set_epi(dat, "nTest", at, nTest)
dat <- set_epi(dat, "nDiagNew", at, nDiagNew)
dat <- set_epi(dat, "nRest", at, length(idsTestedReset))
dat <- set_epi(dat, "nDiag", at, sum(diag.status == 1, na.rm = TRUE))
return(dat)
}- 1
-
Both pairs of attributes (
tested.status/tested.timeanddiag.status/diag.time) are initialized at timestep 2 because timestep 1 is reserved forinit.netsetup. - 2
- Eligible to test: active, not currently in a test window, not currently on an active treatment course.
- 3
-
Each eligible individual independently tests with probability
test.rateper timestep (Bernoulli trial). - 4
-
The key separation. Susceptibles who test receive a negative result — they have
tested.status = 1butdiag.status = 0, so they never enter the treatment pipeline. Only infected testers receivediag.status = 1. - 5
-
Tested-recently flag expires after
test.durtimesteps so the person can be re-tested. - 6
-
If a diagnosed individual has not recovered within
test.durtimesteps, the treatment course ends and they return to the natural clearance rate until re-tested. Recovery itself clearsdiag.statusseparately in the recovery module.
Recovery Module
This module replaces EpiModel’s built-in recovery module to implement heterogeneous recovery rates. Diagnosed individuals recover at rec.rate.tx (fast, antibiotic treatment) while undiagnosed individuals recover at rec.rate (slow, natural clearance). This is the core mechanism of the test-and-treat intervention: screening accelerates recovery for those diagnosed, reducing their infectious duration and thereby suppressing transmission.
recov <- function(dat, at) {
## Attributes ##
active <- get_attr(dat, "active")
status <- get_attr(dat, "status")
diag.status <- get_attr(dat, "diag.status")
## Parameters ##
rec.rate <- get_param(dat, "rec.rate")
rec.rate.tx <- get_param(dat, "rec.rate.tx")
## Determine eligible to recover ##
idsElig <- which(active == 1 & status == "i")
nElig <- length(idsElig)
## Heterogeneous recovery rates based on diagnosis status ##
diag.status.elig <- diag.status[idsElig]
1 ratesElig <- ifelse(diag.status.elig == 1, rec.rate.tx, rec.rate)
## Stochastic recovery process ##
2 vecRecov <- which(rbinom(nElig, 1, ratesElig) == 1)
idsRecov <- idsElig[vecRecov]
nRecov <- length(idsRecov)
## Update attributes for recovered individuals ##
3 status[idsRecov] <- "s"
diag.status[idsRecov] <- 0
## Write out updated attributes ##
dat <- set_attr(dat, "status", status)
dat <- set_attr(dat, "diag.status", diag.status)
## Summary statistics ##
dat <- set_epi(dat, "is.flow", at, nRecov)
return(dat)
}- 1
-
The core intervention mechanism: diagnosed individuals recover at
rec.rate.tx(0.5, mean ~2 weeks) while undiagnosed recover atrec.rate(0.05, mean ~20 weeks) — a 10x difference. - 2
- Each infected individual independently recovers with their diagnosis-specific probability (Bernoulli trial with heterogeneous rates).
- 3
- Recovery clears both disease status (back to susceptible) and diagnosis status, since the individual is cured and exits the treatment pipeline.
Network Model
The formation model uses three ERGM terms to create a realistic sexual contact network for STI transmission.
- 1
-
Three ERGM terms:
edgescontrols mean degree,concurrentcontrols overlapping partnerships (degree > 1), anddegrange(from = 4)prevents unrealistically high-degree nodes. - 2
- Targets: 175 edges (mean degree 0.7), 110 concurrent nodes (22% of the population), and zero nodes with 4+ simultaneous partners.
- 3
- Mean partnership duration of 50 weeks (~1 year).
Dissolution Coefficients
=======================
Dissolution Model: ~offset(edges)
Target Statistics: 50
Crude Coefficient: 3.89182
Mortality/Exit Rate: 0
Adjusted Coefficient: 3.89182Network Diagnostics
dx <- netdx(est, nsims = nsims, ncores = ncores, nsteps = nsteps,
nwstats.formula = ~edges + concurrent + degree(0:5))
Network Diagnostics
-----------------------
- Simulating 5 networks
- Calculating formation statisticsprint(dx)EpiModel Network Diagnostics
=======================
Diagnostic Method: Dynamic
Simulations: 5
Time Steps per Sim: 500
Formation Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 175 173.987 -0.579 1.763 -0.575 3.887 11.824
concurrent 110 108.066 -1.758 1.268 -1.525 2.953 11.057
degree0 NA 273.404 NA 2.230 NA 4.127 13.063
degree1 NA 118.529 NA 1.080 NA 1.884 10.339
degree2 NA 94.754 NA 1.102 NA 2.163 9.969
degree3 NA 13.312 NA 0.365 NA 0.887 3.626
degree4 NA 0.000 NA NaN NA 0.000 0.000
degree5 NA 0.000 NA NaN NA 0.000 0.000
Duration Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 50 49.499 -1.002 0.484 -1.035 0.617 3.202
Dissolution Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.02 0.02 0.918 0 0.891 0 0.011plot(dx)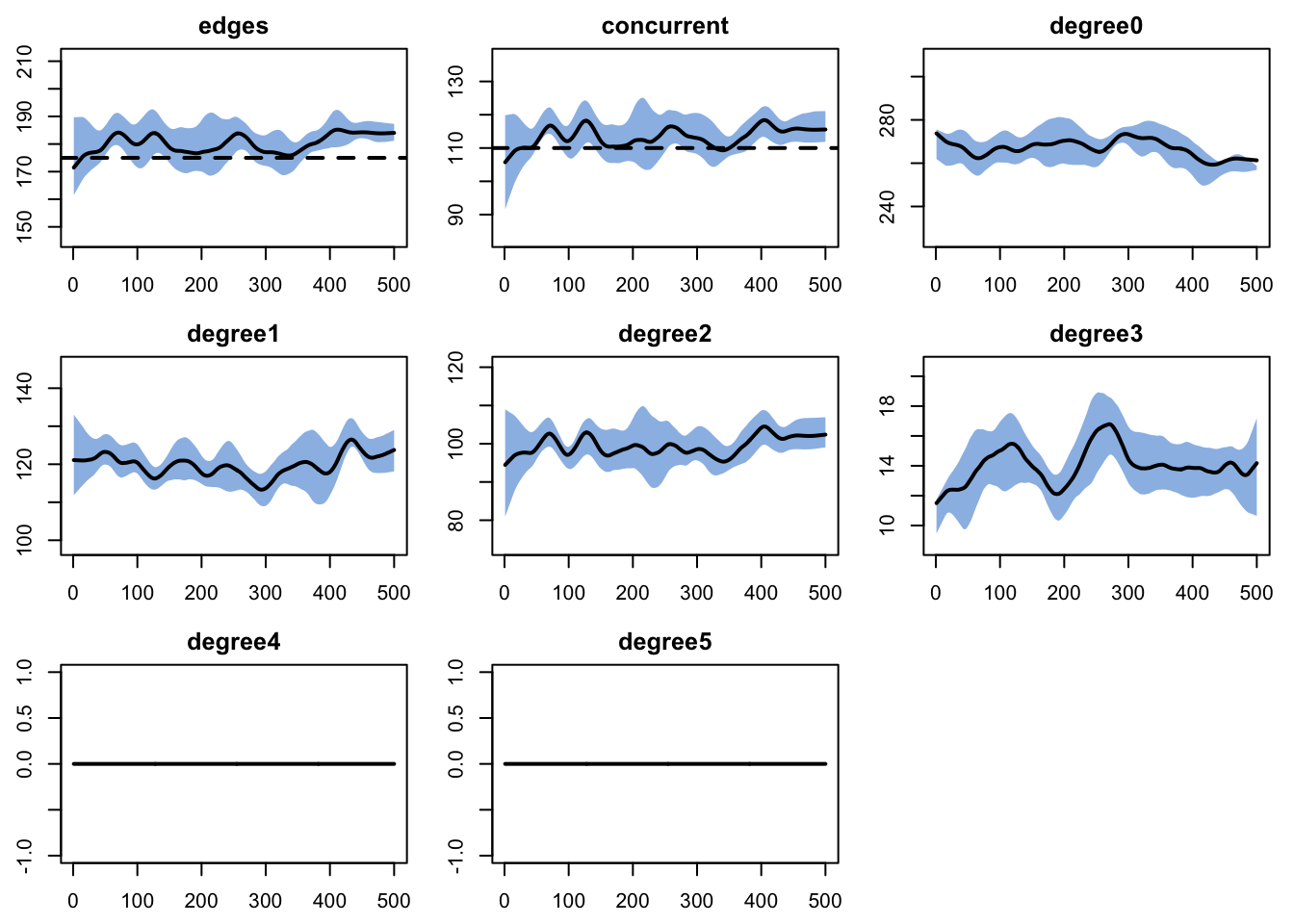
Epidemic Simulation
Initial Conditions and Control Settings
All three scenarios share the same initial conditions and control settings. The control.net call registers the built-in infection module alongside the two custom modules (test-and-treat and recovery).
- 1
- Seed the epidemic with 10 infected nodes (2% of the 500-node population).
- 2
-
type = NULLtells EpiModel that all modules are explicitly specified — no built-in SIS/SIR logic. - 3
-
infection.netis EpiModel’s built-in SI infection module;recovandtntare the custom modules defined above.
Scenario 1: No Screening (SIS Baseline)
Without any screening, only natural clearance (rec.rate = 0.05) drives recovery. The SIS model reaches an endemic equilibrium where new infections balance natural recoveries. This is the counterfactual — what happens without intervention.
param_none <- param.net(
inf.prob = 0.4, act.rate = 2,
rec.rate = 1 / 20, rec.rate.tx = 0.5,
test.rate = 0, test.dur = 2
)
sim_none <- netsim(est, param_none, init, control)
print(sim_none)EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type:
No. simulations: 5
No. time steps: 500
No. NW groups: 1
Fixed Parameters
---------------------------
inf.prob = 0.4
act.rate = 2
rec.rate = 0.05
rec.rate.tx = 0.5
test.rate = 0
test.dur = 2
groups = 1
Model Functions
-----------------------
initialize.FUN
resim_nets.FUN
summary_nets.FUN
infection.FUN
recovery.FUN
nwupdate.FUN
prevalence.FUN
verbose.FUN
tnt.FUN
Model Output
-----------------------
Variables: s.num i.num num nTest nDiagNew nRest nDiag
si.flow is.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 175 174.629 -0.212 1.633 -0.227 6.103 11.523
concurrent 110 109.022 -0.889 1.087 -0.899 4.343 10.368
deg4+ 0 0.000 NaN NaN NaN 0.000 0.000
Duration Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 50 49.864 -0.271 0.709 -0.191 1.185 3.92
Dissolution Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.02 0.02 -1.596 0 -1.493 0 0.011Scenario 2: Standard Screening
10% of undiagnosed individuals test each week (~10-week average interval between tests). This represents a moderate screening program — feasible for clinical settings with annual or semi-annual testing recommendations.
param_std <- param.net(
inf.prob = 0.4, act.rate = 2,
rec.rate = 1 / 20, rec.rate.tx = 0.5,
test.rate = 0.1, test.dur = 2
)
sim_std <- netsim(est, param_std, init, control)
print(sim_std)EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type:
No. simulations: 5
No. time steps: 500
No. NW groups: 1
Fixed Parameters
---------------------------
inf.prob = 0.4
act.rate = 2
rec.rate = 0.05
rec.rate.tx = 0.5
test.rate = 0.1
test.dur = 2
groups = 1
Model Functions
-----------------------
initialize.FUN
resim_nets.FUN
summary_nets.FUN
infection.FUN
recovery.FUN
nwupdate.FUN
prevalence.FUN
verbose.FUN
tnt.FUN
Model Output
-----------------------
Variables: s.num i.num num nTest nDiagNew nRest nDiag
si.flow is.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 175 175.416 0.238 3.035 0.137 8.490 17.121
concurrent 110 110.056 0.051 1.950 0.029 7.046 14.751
deg4+ 0 0.000 NaN NaN NaN 0.000 0.000
Duration Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 50 50.194 0.389 0.532 0.365 1.657 3.453
Dissolution Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.02 0.02 -1.825 0 -1.889 0 0.01Scenario 3: Intensive Screening
30% of undiagnosed individuals test each week (~3-week average interval). This represents an aggressive public health intervention — frequent screening in high-prevalence populations.
param_int <- param.net(
inf.prob = 0.4, act.rate = 2,
rec.rate = 1 / 20, rec.rate.tx = 0.5,
test.rate = 0.3, test.dur = 2
)
sim_int <- netsim(est, param_int, init, control)
print(sim_int)EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type:
No. simulations: 5
No. time steps: 500
No. NW groups: 1
Fixed Parameters
---------------------------
inf.prob = 0.4
act.rate = 2
rec.rate = 0.05
rec.rate.tx = 0.5
test.rate = 0.3
test.dur = 2
groups = 1
Model Functions
-----------------------
initialize.FUN
resim_nets.FUN
summary_nets.FUN
infection.FUN
recovery.FUN
nwupdate.FUN
prevalence.FUN
verbose.FUN
tnt.FUN
Model Output
-----------------------
Variables: s.num i.num num nTest nDiagNew nRest nDiag
si.flow is.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 175 175.884 0.505 2.147 0.412 6.735 13.327
concurrent 110 109.136 -0.785 1.588 -0.544 6.470 12.741
deg4+ 0 0.000 NaN NaN NaN 0.000 0.000
Duration Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 50 49.731 -0.539 0.513 -0.525 2.137 3.588
Dissolution Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.02 0.02 0.179 0 0.17 0 0.011Analysis
sim_none <- mutate_epi(sim_none, prev = i.num / num)
sim_std <- mutate_epi(sim_std, prev = i.num / num)
sim_int <- mutate_epi(sim_int, prev = i.num / num)Prevalence Comparison
The central question: how does screening intensity affect population-level STI prevalence? Without screening, the model reaches an endemic equilibrium at roughly 45%. Standard screening reduces prevalence to approximately 30%, and intensive screening pushes it to near elimination.
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_none, y = "prev",
main = "Prevalence by Screening Intensity",
ylab = "Prevalence (I / N)", xlab = "Week",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim_std, y = "prev",
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE, add = TRUE)
plot(sim_int, y = "prev",
mean.col = "seagreen", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "seagreen", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE, add = TRUE)
legend("topright",
legend = c("No Screening", "Standard (10%/wk)", "Intensive (30%/wk)"),
col = c("firebrick", "steelblue", "seagreen"), lwd = 2, bty = "n")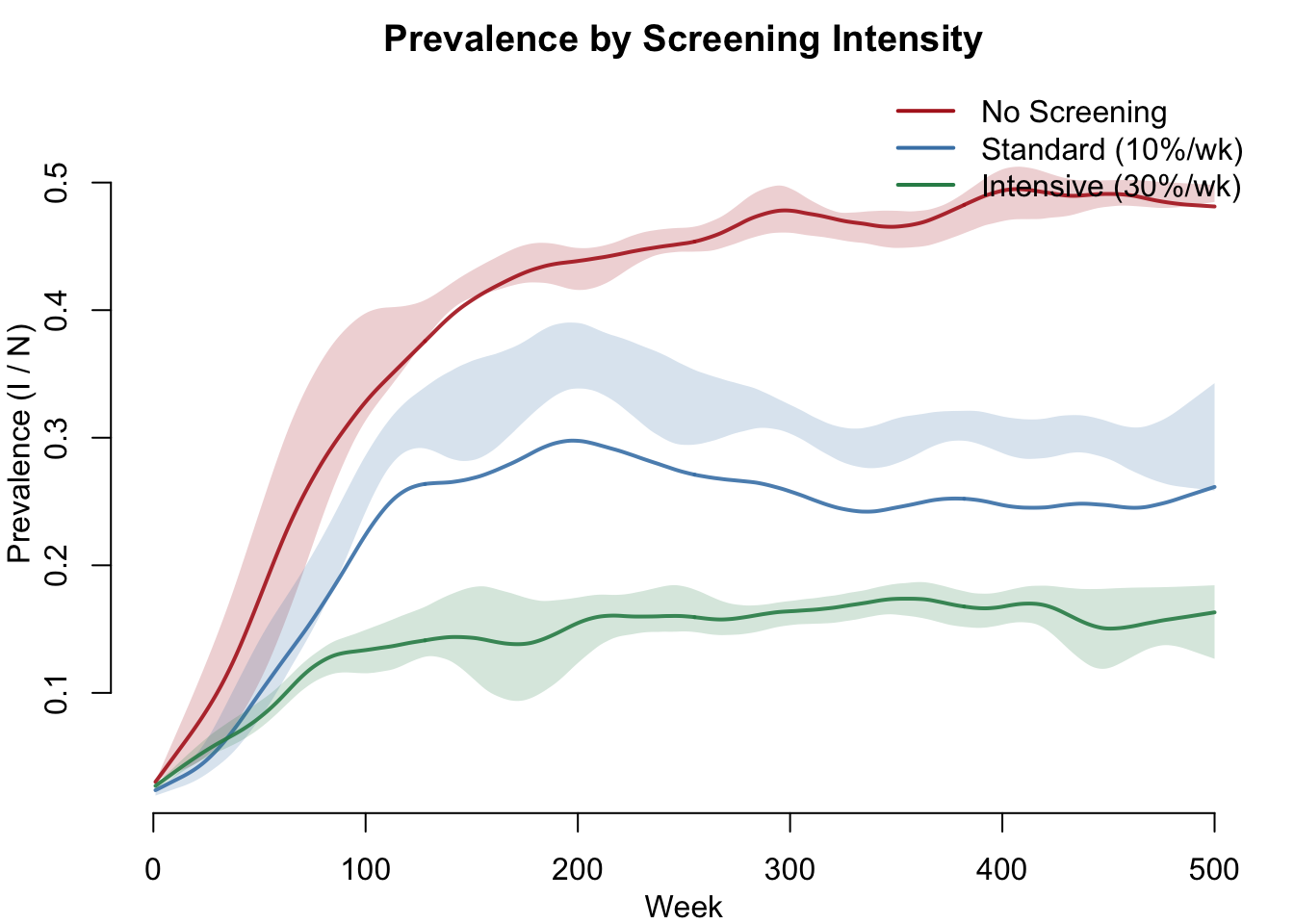
Compartment Dynamics
A side-by-side comparison of susceptible and infectious compartment counts. Without screening, the SIS model reaches a stable endemic equilibrium with a large infected pool. Intensive screening suppresses the infectious compartment dramatically.
par(mfrow = c(1, 2), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_none, y = c("s.num", "i.num"),
main = "No Screening",
mean.col = c("steelblue", "firebrick"), mean.lwd = 2, mean.smooth = TRUE,
qnts = FALSE, legend = TRUE)
plot(sim_int, y = c("s.num", "i.num"),
main = "Intensive Screening",
mean.col = c("steelblue", "firebrick"), mean.lwd = 2, mean.smooth = TRUE,
qnts = FALSE, legend = TRUE)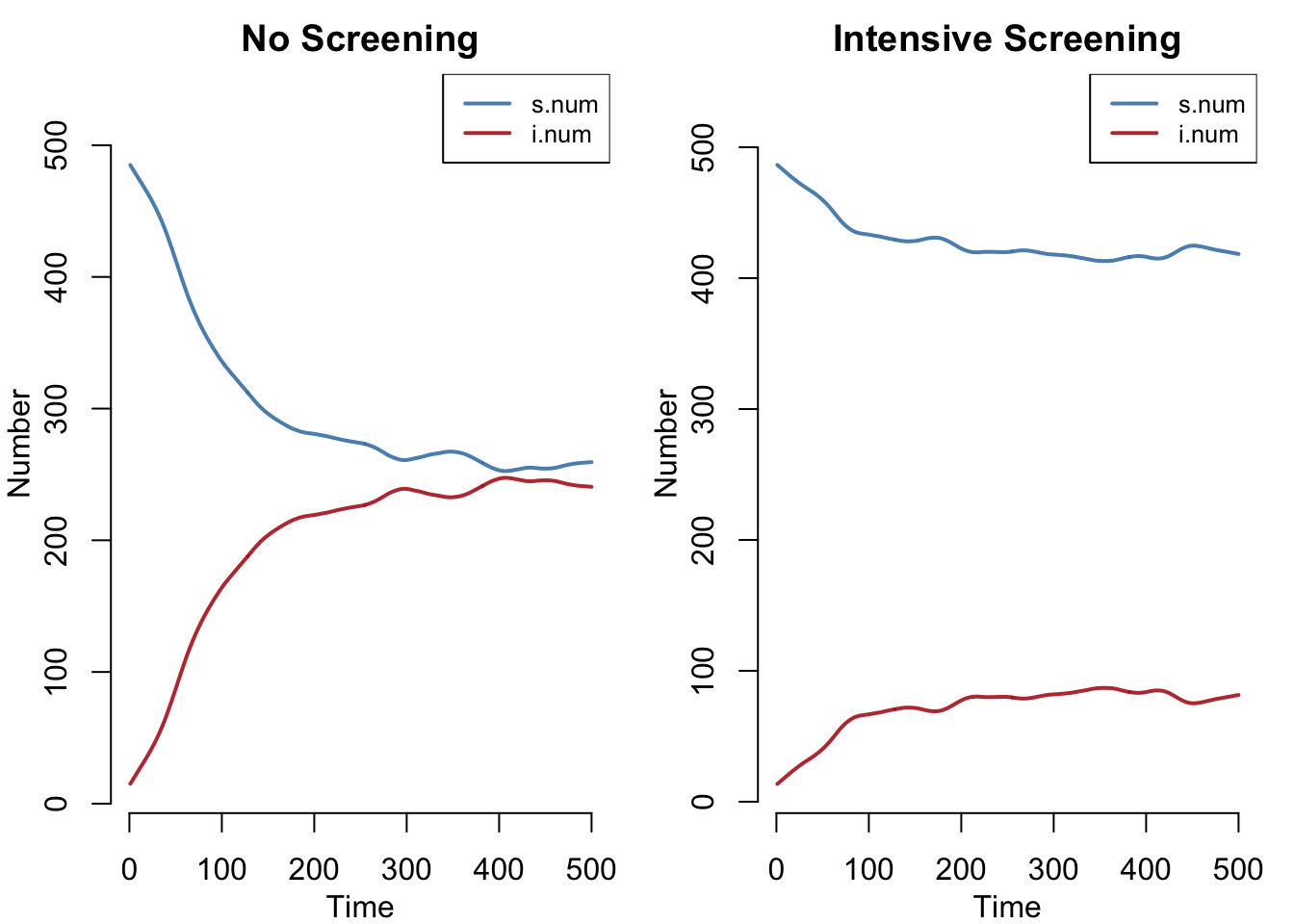
Diagnosis Coverage
How many individuals are currently diagnosed at each timestep? This shows the test-and-treat cascade in action — the fraction of infected individuals who are in the treatment pipeline.
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_std, y = "nDiag",
main = "Currently Diagnosed (Treatment Pipeline)",
ylab = "Number Diagnosed", xlab = "Week",
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim_int, y = "nDiag",
mean.col = "#27ae60", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "#27ae60", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE, add = TRUE)
legend("topright",
legend = c("Standard (10%/wk)", "Intensive (30%/wk)"),
col = c("steelblue", "#27ae60"), lwd = 2, bty = "n")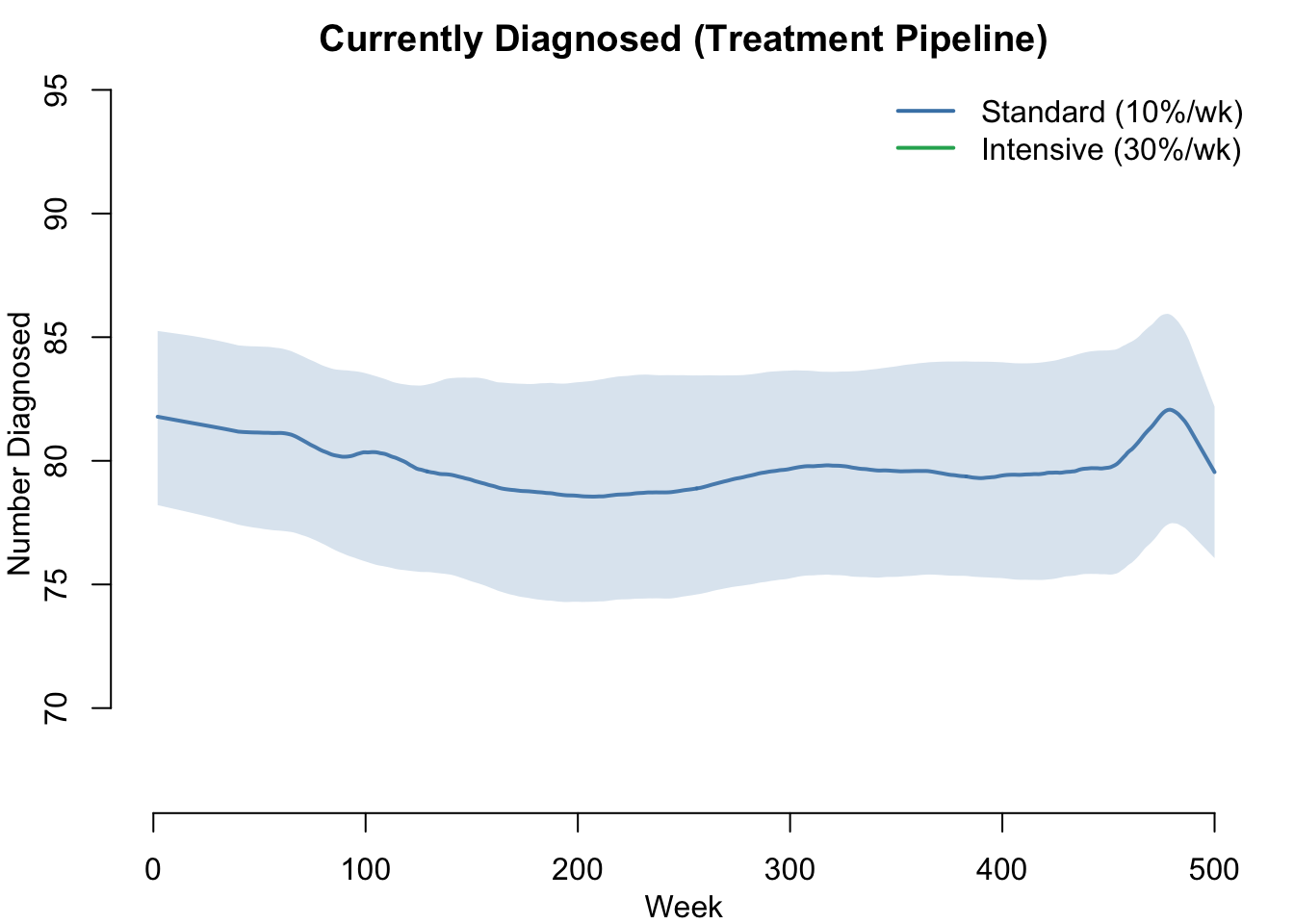
Incidence
Incidence (new infections per week) shows the epidemic’s momentum. Without screening, incidence stabilizes at a high endemic level. With screening, incidence drops as fewer infected individuals are available to transmit.
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_none, y = "si.flow",
main = "New Infections per Week",
ylab = "New Infections", xlab = "Week",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim_std, y = "si.flow",
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE, add = TRUE)
plot(sim_int, y = "si.flow",
mean.col = "#27ae60", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "#27ae60", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE, add = TRUE)
legend("topright",
legend = c("No Screening", "Standard (10%/wk)", "Intensive (30%/wk)"),
col = c("firebrick", "steelblue", "#27ae60"), lwd = 2, bty = "n")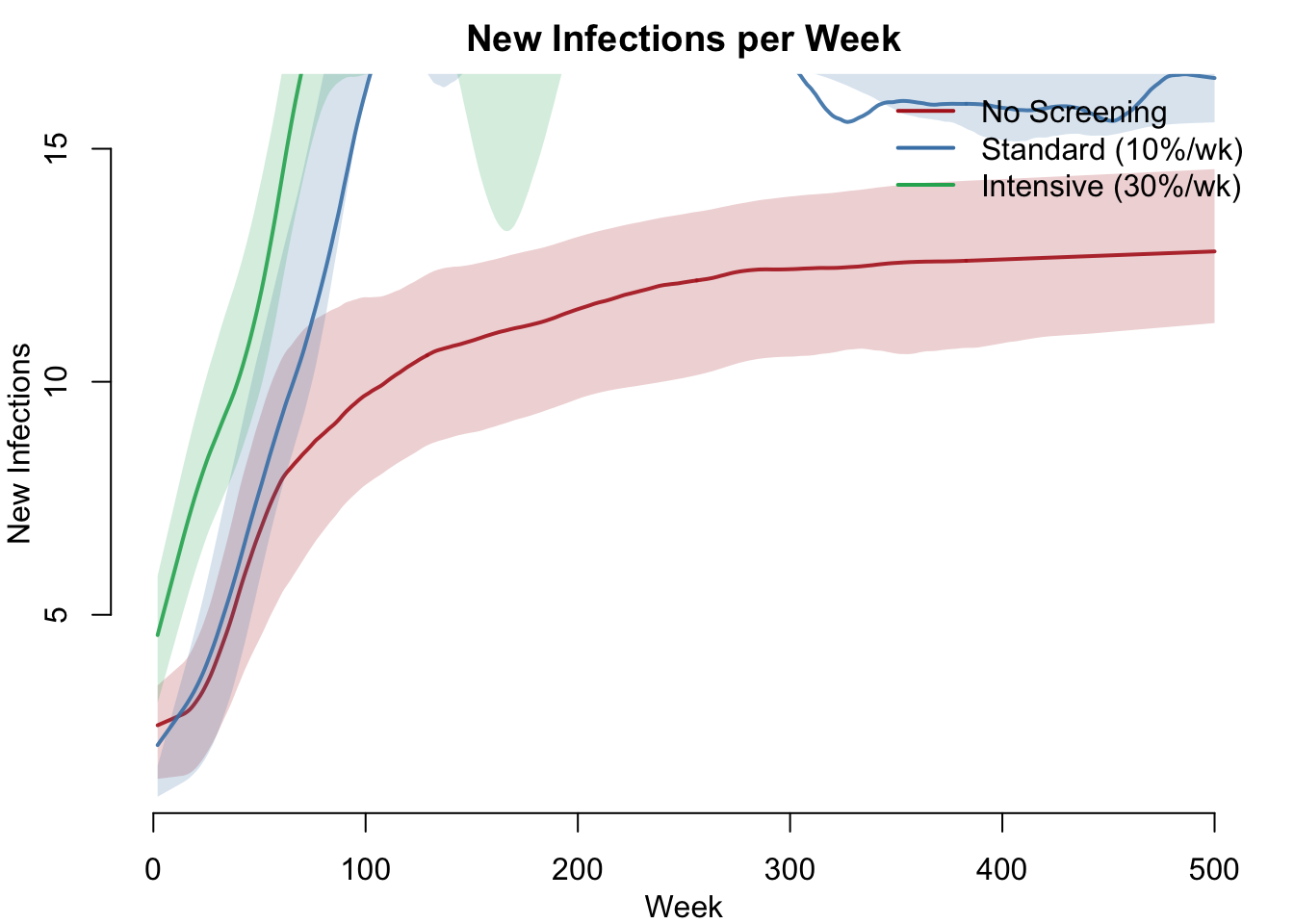
Summary Table
In an SIS model, the right burden metric is person-weeks infected (total time spent in the I compartment across all individuals), not cumulative new infections. Cumulative infections can paradoxically increase with screening because treated individuals recover faster, become susceptible again, and get reinfected.
df_none <- as.data.frame(sim_none)
df_std <- as.data.frame(sim_std)
df_int <- as.data.frame(sim_int)
pw_none <- mean(tapply(df_none$i.num, df_none$sim, sum, na.rm = TRUE))
pw_std <- mean(tapply(df_std$i.num, df_std$sim, sum, na.rm = TRUE))
pw_int <- mean(tapply(df_int$i.num, df_int$sim, sum, na.rm = TRUE))
knitr::kable(data.frame(
Metric = c("Equilibrium prevalence",
"Person-weeks infected",
"Burden reduction (vs. none)",
"Mean diagnosed (nDiag)"),
No_Screening = c(
round(mean(df_none$prev[df_none$time > nsteps * 0.8], na.rm = TRUE), 3),
round(pw_none),
"---",
0
),
Standard = c(
round(mean(df_std$prev[df_std$time > nsteps * 0.8], na.rm = TRUE), 3),
round(pw_std),
paste0(round((1 - pw_std / pw_none) * 100), "%"),
round(mean(df_std$nDiag[df_std$time > nsteps * 0.8], na.rm = TRUE), 1)
),
Intensive = c(
round(mean(df_int$prev[df_int$time > nsteps * 0.8], na.rm = TRUE), 3),
round(pw_int),
paste0(round((1 - pw_int / pw_none) * 100), "%"),
round(mean(df_int$nDiag[df_int$time > nsteps * 0.8], na.rm = TRUE), 1)
)
))| Metric | No_Screening | Standard | Intensive |
|---|---|---|---|
| Equilibrium prevalence | 0.447 | 0.375 | 0.244 |
| Person-weeks infected | 99604 | 76018 | 53403 |
| Burden reduction (vs. none) | — | 24% | 46% |
| Mean diagnosed (nDiag) | 0 | 25 | 40.3 |
Sensitivity Analysis: Prevalence vs. Testing Rate
How does equilibrium prevalence respond to increasing screening intensity? This dose-response curve sweeps test.rate from 0 to 0.5, showing the marginal return of additional testing effort and helping identify the optimal screening intensity for intervention design.
test_rates <- seq(0, 0.5, by = 0.05)
eq_prev <- numeric(length(test_rates))
for (i in seq_along(test_rates)) {
param_i <- param.net(
inf.prob = 0.4, act.rate = 2,
rec.rate = 1 / 20, rec.rate.tx = 0.5,
test.rate = test_rates[i], test.dur = 2
)
sim_i <- netsim(est, param_i, init, control)
sim_i <- mutate_epi(sim_i, prev = i.num / num)
df_i <- as.data.frame(sim_i)
eq_prev[i] <- mean(df_i$prev[df_i$time > nsteps * 0.8], na.rm = TRUE)
}
# Indices for the three named scenarios
idx_none <- which.min(abs(test_rates - 0))
idx_std <- which.min(abs(test_rates - 0.1))
idx_int <- which.min(abs(test_rates - 0.3))
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(test_rates, eq_prev, type = "b", pch = 19, col = "steelblue",
lwd = 2, xlab = "Weekly Testing Rate",
ylab = "Equilibrium Prevalence",
main = "Prevalence vs. Screening Intensity",
ylim = c(0, max(eq_prev) * 1.15))
abline(h = 0, lty = 2, col = "gray50")
points(test_rates[c(idx_none, idx_std, idx_int)],
eq_prev[c(idx_none, idx_std, idx_int)],
pch = 19, col = "firebrick", cex = 1.5)
text(0.05, eq_prev[idx_none] + 0.02, "No screening", col = "firebrick", cex = 0.8)
text(0.15, eq_prev[idx_std] + 0.02, "Standard", col = "firebrick", cex = 0.8)
text(0.35, eq_prev[idx_int] + 0.02, "Intensive", col = "firebrick", cex = 0.8)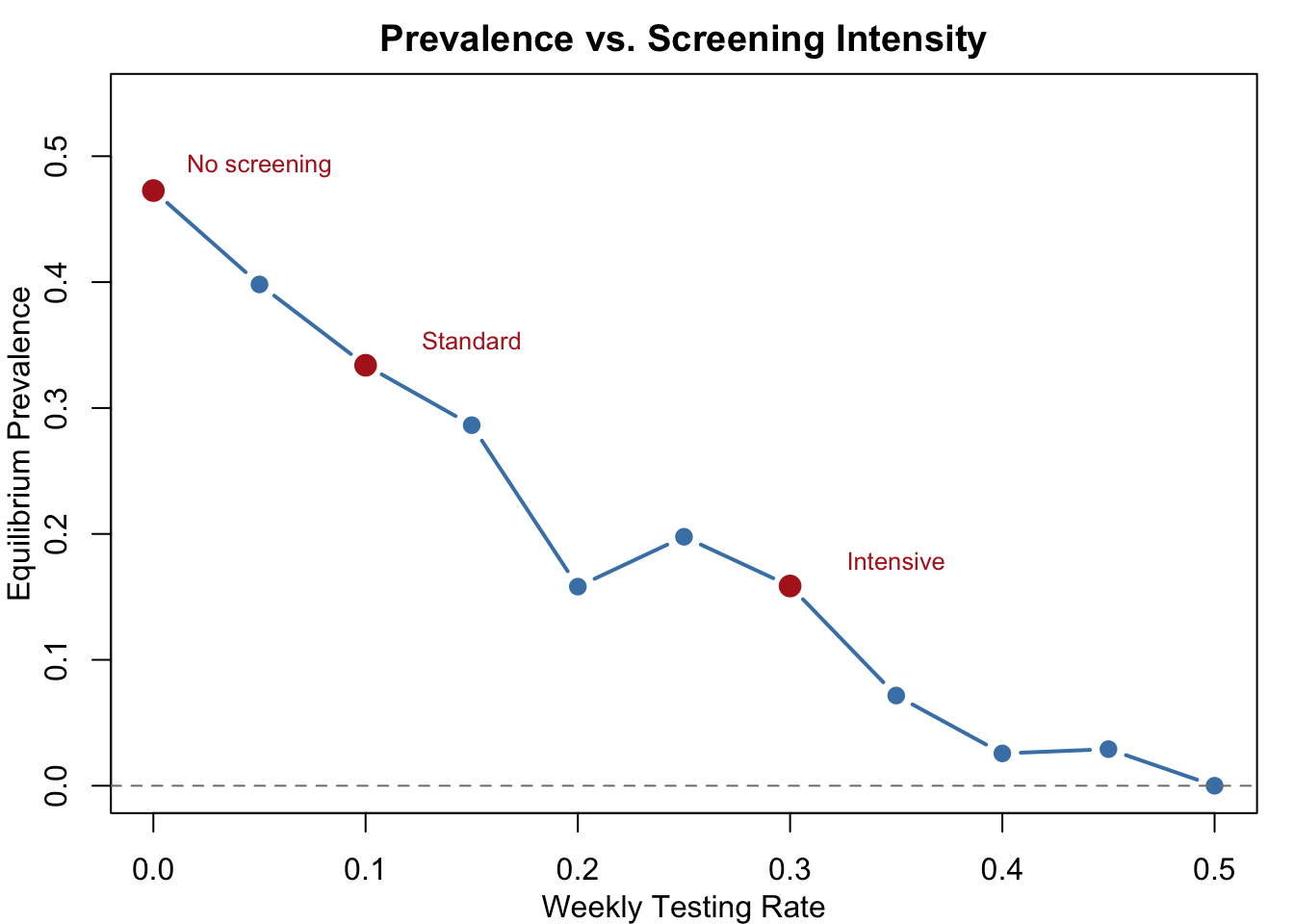
knitr::kable(data.frame(
test.rate = test_rates,
eq.prevalence = round(eq_prev, 3)
))| test.rate | eq.prevalence |
|---|---|
| 0.00 | 0.445 |
| 0.05 | 0.396 |
| 0.10 | 0.329 |
| 0.15 | 0.319 |
| 0.20 | 0.278 |
| 0.25 | 0.262 |
| 0.30 | 0.232 |
| 0.35 | 0.217 |
| 0.40 | 0.210 |
| 0.45 | 0.197 |
| 0.50 | 0.183 |
Parameters
Transmission
| Parameter | Description | Default |
|---|---|---|
inf.prob |
Per-act transmission probability | 0.4 |
act.rate |
Acts per partnership per timestep | 2 |
Recovery and Treatment
| Parameter | Description | Default |
|---|---|---|
rec.rate |
Natural clearance rate (undiagnosed) | 1/20 (mean ~20 weeks) |
rec.rate.tx |
Treatment recovery rate (diagnosed) | 0.5 (mean ~2 weeks) |
Screening
| Parameter | Description | Varied |
|---|---|---|
test.rate |
Weekly testing rate for undiagnosed individuals | 0 / 0.1 / 0.3 |
test.dur |
Duration diagnosis persists (timesteps) | 2 |
Network
| Parameter | Description | Default |
|---|---|---|
| Population size | Number of nodes | 500 |
| Target edges | Mean concurrent partnerships | 175 |
| Concurrency | Nodes with degree > 1 | 110 |
| Max degree | Upper bound on simultaneous partners | 3 |
| Partnership duration | Mean edge duration (weeks) | 50 |
Module Execution Order
resim_nets → infection → tnt → recovery → prevalenceThe tnt module runs after infection (so newly infected individuals can be tested in the same timestep) and before recovery (so newly diagnosed individuals immediately benefit from the higher treatment recovery rate).
Next Steps
- Separate testing from treatment: Add a treatment uptake probability or delay between diagnosis and treatment initiation — not all diagnosed individuals start treatment immediately
- Add partner notification / contact tracing: Diagnosed individuals’ partners receive expedited testing, amplifying the intervention’s reach
- Risk-based screening: Vary
test.rateby individual attributes (e.g., higher-risk groups test more frequently) - Add vital dynamics for longer-term STI modeling — see SI with Vital Dynamics
- Model antimicrobial resistance: Introduce treatment failure or competing resistant strains — see Competing Strains
- Extend to multi-stage disease: Model progression through disease stages with stage-dependent testing and treatment — see Syphilis
- Add cost-effectiveness analysis: Attach costs to testing and treatment to evaluate intervention efficiency — see Cost-Effectiveness Analysis