flowchart LR
in(( )) -->|"arrivals"| S
S["<b>S</b><br/>Susceptible"] -->|"infection"| I1["<b>Acute</b><br/>10x inf."]
I1 -->|"progression"| I2["<b>Chronic 1</b><br/>1x inf."]
I2 -->|"progression"| I3["<b>Chronic 2</b><br/>1x inf."]
I3 -->|"progression"| I4["<b>AIDS</b><br/>5x inf."]
S -->|"departure"| out1(( ))
I1 -->|"departure"| out2(( ))
I2 -->|"departure"| out3(( ))
I3 -->|"departure"| out4(( ))
I4 -->|"elevated<br/>departure"| out5(( ))
style S fill:#3498db,color:#fff
style I1 fill:#e67e22,color:#fff
style I2 fill:#e74c3c,color:#fff
style I3 fill:#c0392b,color:#fff
style I4 fill:#8e44ad,color:#fff
style in fill:none,stroke:none
style out1 fill:none,stroke:none
style out2 fill:none,stroke:none
style out3 fill:none,stroke:none
style out4 fill:none,stroke:none
style out5 fill:none,stroke:none
HIV Transmission Model
Overview
This example implements a simplified HIV transmission model over a dynamic network, based on the deterministic compartmental framework from Granich et al. (2009). The model extends EpiModel’s built-in SI framework with four distinct infectious sub-compartments — Acute, Chronic 1, Chronic 2, and AIDS — each with different levels of infectiousness. On top of this disease progression structure, the model layers an antiretroviral therapy (ART) intervention that reduces both infectiousness and the rate of disease progression.
The central question, following Granich et al., is whether universal ART — treating all HIV-positive individuals regardless of disease stage — can reduce population-level HIV prevalence and incidence. The model compares two scenarios: one with ART (treatment rate of 1% per week among untreated infected individuals) and one without ART, allowing us to quantify the prevention benefit of treatment-as-prevention.
This is one of the more advanced examples in the Gallery. It demonstrates how to build a fully custom module set (type = NULL) with multiple interacting processes: stage-dependent transmission, ART dynamics, disease progression with treatment modification, and differential mortality. The patterns here — multiplicative transmission probability, layered treatment states, and multiple flow trackers — are building blocks for more complex HIV models with care cascades, PrEP, or demographic heterogeneity.
- model.R — Main simulation script
- module-fx.R — Custom module functions
Model Structure
Disease Compartments
| Compartment | Label | Description |
|---|---|---|
| Susceptible | S | Not infected with HIV |
| Acute | Acute | Recently infected; high viral load produces elevated infectiousness |
| Chronic 1 | Chronic 1 | Early chronic infection; low infectiousness, stable CD4+ count |
| Chronic 2 | Chronic 2 | Late chronic infection; low infectiousness, declining CD4+ count |
| AIDS | AIDS | Advanced disease; rising viral load increases infectiousness, elevated mortality |
Each infectious compartment has two sub-states: on ART and not on ART. ART reduces infectiousness by 95% and halves the rate of disease progression and AIDS mortality.
Transmission Probability
Transmission probability follows a multiplicative structure. The base per-act probability during chronic infection is scaled by the infector’s disease stage and ART status:
| Infector Stage | Multiplier | Default |
|---|---|---|
| Acute | relative.inf.prob.acute |
10x |
| Chronic 1 or 2 | 1 (reference) | 1x |
| AIDS | relative.inf.prob.AIDS |
5x |
| ART Status | Multiplier | Default |
|---|---|---|
| Not on ART | 1 | 1x |
| On ART | relative.inf.prob.ART |
0.05x |
The per-act probability is then converted to a per-timestep probability accounting for multiple acts per partnership: finalProb = 1 - (1 - transProb)^act.rate.
Setup
suppressMessages(library(EpiModel))
nsims <- 5
ncores <- 5
nsteps <- 500Custom Modules
Infection Module
Simulates HIV transmission from infected to susceptible individuals on discordant edges. The transmission probability depends on the infector’s disease stage and ART status, using the multiplicative structure described above. Newly infected individuals enter the acute stage with no ART.
infect <- function(dat, at) {
## Attributes ##
active <- get_attr(dat, "active")
status <- get_attr(dat, "status")
ART.status <- get_attr(dat, "ART.status")
stage <- get_attr(dat, "stage")
stage.time <- get_attr(dat, "stage.time")
ART.time <- get_attr(dat, "ART.time")
## Parameters ##
inf.prob.chronic <- get_param(dat, "inf.prob.chronic")
relative.inf.prob.acute <- get_param(dat, "relative.inf.prob.acute")
relative.inf.prob.AIDS <- get_param(dat, "relative.inf.prob.AIDS")
relative.inf.prob.ART <- get_param(dat, "relative.inf.prob.ART")
act.rate <- get_param(dat, "act.rate")
## Find infected nodes ##
idsInf <- which(active == 1 & status == "i")
nActive <- sum(active == 1)
nElig <- length(idsInf)
## Initialize default incidence at 0 ##
nInf <- 0
## If any infected nodes, proceed with transmission ##
if (nElig > 0 && nElig < nActive) {
## Look up discordant edgelist ##
del <- discord_edgelist(dat, at)
## If any discordant pairs, proceed ##
if (!is.null(del)) {
stage_mult <- ifelse(stage[del$inf] == "acute",
relative.inf.prob.acute,
ifelse(stage[del$inf] == "AIDS",
1 relative.inf.prob.AIDS, 1))
art_mult <- ifelse(ART.status[del$inf] == 1,
2 relative.inf.prob.ART, 1)
del$transProb <- inf.prob.chronic * stage_mult * art_mult
del$actRate <- act.rate
3 del$finalProb <- 1 - (1 - del$transProb)^del$actRate
# Stochastic transmission process
transmit <- rbinom(nrow(del), 1, del$finalProb)
del <- del[which(transmit == 1), ]
idsNewInf <- unique(del$sus)
nInf <- length(idsNewInf)
# Set new attributes for those newly infected
if (nInf > 0) {
status[idsNewInf] <- "i"
stage[idsNewInf] <- "acute"
ART.status[idsNewInf] <- 0
stage.time[idsNewInf] <- 0
4 ART.time[idsNewInf] <- 0
}
}
}
## Update attributes ##
dat <- set_attr(dat, "status", status)
dat <- set_attr(dat, "stage", stage)
dat <- set_attr(dat, "ART.status", ART.status)
dat <- set_attr(dat, "stage.time", stage.time)
dat <- set_attr(dat, "ART.time", ART.time)
## Save summary statistics ##
dat <- set_epi(dat, "acute.flow", at, nInf)
dat <- set_epi(dat, "s.num", at, sum(active == 1 & status == "s"))
dat <- set_epi(dat, "acute.ART.num", at,
sum(active == 1 & stage == "acute" &
ART.status == 1, na.rm = TRUE))
dat <- set_epi(dat, "acute.NoART.num", at,
sum(active == 1 & stage == "acute" &
ART.status == 0, na.rm = TRUE))
dat <- set_epi(dat, "chronic1.ART.num", at,
sum(active == 1 & stage == "chronic1" &
ART.status == 1, na.rm = TRUE))
dat <- set_epi(dat, "chronic1.NoART.num", at,
sum(active == 1 & stage == "chronic1" &
ART.status == 0, na.rm = TRUE))
dat <- set_epi(dat, "chronic2.ART.num", at,
sum(active == 1 & stage == "chronic2" &
ART.status == 1, na.rm = TRUE))
dat <- set_epi(dat, "chronic2.NoART.num", at,
sum(active == 1 & stage == "chronic2" &
ART.status == 0, na.rm = TRUE))
dat <- set_epi(dat, "AIDS.ART.num", at,
sum(active == 1 & stage == "AIDS" &
ART.status == 1, na.rm = TRUE))
dat <- set_epi(dat, "AIDS.NoART.num", at,
sum(active == 1 & stage == "AIDS" &
5 ART.status == 0, na.rm = TRUE))
return(dat)
}- 1
- Stage multiplier: acute-stage infectors are 10x more infectious than chronic; AIDS-stage infectors are 5x.
- 2
- ART multiplier: individuals on ART have 95% reduced infectiousness (0.05x).
- 3
-
Per-timestep probability from multiple acts: accounts for
act.rateindependent transmission opportunities per partnership per week. - 4
- Newly infected individuals enter as acute, not on ART, with time counters set to 0 to prevent same-step transitions.
- 5
- Track counts for each stage-by-ART combination to enable detailed analysis of the epidemic composition.
Progression Module
Handles two interrelated processes: ART dynamics (treatment initiation and discontinuance) and disease stage transitions. ART reduces all progression rates by ART.Progression.Reduction.Rate. Both processes use a one-timestep delay — individuals cannot transition out of a state in the same timestep they entered it.
progress <- function(dat, at) {
## Attributes ##
active <- get_attr(dat, "active")
status <- get_attr(dat, "status")
## Initialize stage/ART tracking attributes at simulation start ##
if (at == 2) {
dat <- set_attr(dat, "stage",
ifelse(status == "i", "acute", NA))
dat <- set_attr(dat, "ART.status",
ifelse(status == "i", 0, NA))
dat <- set_attr(dat, "stage.time",
ifelse(status == "i", 0, NA))
dat <- set_attr(dat, "ART.time",
1 ifelse(status == "i", 0, NA))
}
## Attributes (read after initialization) ##
ART.status <- get_attr(dat, "ART.status")
stage <- get_attr(dat, "stage")
ART.time <- get_attr(dat, "ART.time")
stage.time <- get_attr(dat, "stage.time")
## Parameters ##
AcuteToChronic1.Rate <- get_param(dat, "AcuteToChronic1.Rate")
Chronic1ToChronic2.Rate <- get_param(dat, "Chronic1ToChronic2.Rate")
Chronic2ToAIDS.Rate <- get_param(dat, "Chronic2ToAIDS.Rate")
ART.Treatment.Rate <- get_param(dat, "ART.Treatment.Rate")
ART.Discontinuance.Rate <- get_param(dat, "ART.Discontinuance.Rate")
ART.Progression.Reduction.Rate <- get_param(dat,
"ART.Progression.Reduction.Rate")
## Increment time-in-stage and time-on/off-ART counters ##
ART.time <- ifelse(!is.na(ART.time), ART.time + 1, ART.time)
stage.time <- ifelse(!is.na(stage.time),
2 stage.time + 1, stage.time)
## ---- ART Treatment ---- ##
idsEligART <- which(active == 1 & status == "i" &
ART.status == 0 & ART.time != 0 &
!is.na(ART.status))
idsART <- idsEligART[rbinom(length(idsEligART), 1,
ART.Treatment.Rate) == 1]
if (length(idsART) > 0) {
ART.status[idsART] <- 1
ART.time[idsART] <- 0
}
## ---- ART Discontinuance ---- ##
idsEligARTDisc <- which(active == 1 & status == "i" &
ART.status == 1 & ART.time != 0 &
!is.na(ART.status))
idsARTDisc <- idsEligARTDisc[rbinom(length(idsEligARTDisc), 1,
ART.Discontinuance.Rate) == 1]
if (length(idsARTDisc) > 0) {
ART.status[idsARTDisc] <- 0
3 ART.time[idsARTDisc] <- 0
}
## Save ART flow statistics ##
dat <- set_epi(dat, "acute.ART.treatment.flow", at,
sum(stage[idsART] == "acute"))
dat <- set_epi(dat, "acute.ART.discontinuance.flow", at,
sum(stage[idsARTDisc] == "acute"))
dat <- set_epi(dat, "chronic1.ART.treatment.flow", at,
sum(stage[idsART] == "chronic1"))
dat <- set_epi(dat, "chronic1.ART.discont.flow", at,
sum(stage[idsARTDisc] == "chronic1"))
dat <- set_epi(dat, "chronic2.ART.treatment.flow", at,
sum(stage[idsART] == "chronic2"))
dat <- set_epi(dat, "chronic2.ART.discont.flow", at,
sum(stage[idsARTDisc] == "chronic2"))
dat <- set_epi(dat, "AIDS.ART.treatment.flow", at,
sum(stage[idsART] == "AIDS"))
dat <- set_epi(dat, "AIDS.ART.discontinuance.flow", at,
sum(stage[idsARTDisc] == "AIDS"))
## ---- Acute -> Chronic 1 Progression ---- ##
idsEligC1 <- which(active == 1 & stage == "acute" &
stage.time != 0 & !is.na(ART.status))
prog.rate.C1 <- ifelse(ART.status[idsEligC1] == 1,
AcuteToChronic1.Rate *
ART.Progression.Reduction.Rate,
4 AcuteToChronic1.Rate)
idsChronic1 <- idsEligC1[rbinom(length(idsEligC1), 1,
prog.rate.C1) == 1]
if (length(idsChronic1) > 0) {
stage[idsChronic1] <- "chronic1"
stage.time[idsChronic1] <- 0
}
## ---- Chronic 1 -> Chronic 2 Progression ---- ##
idsEligC2 <- which(active == 1 & stage == "chronic1" &
stage.time != 0 & !is.na(ART.status))
prog.rate.C2 <- ifelse(ART.status[idsEligC2] == 1,
Chronic1ToChronic2.Rate *
ART.Progression.Reduction.Rate,
Chronic1ToChronic2.Rate)
idsChronic2 <- idsEligC2[rbinom(length(idsEligC2), 1,
prog.rate.C2) == 1]
if (length(idsChronic2) > 0) {
stage[idsChronic2] <- "chronic2"
stage.time[idsChronic2] <- 0
}
## ---- Chronic 2 -> AIDS Progression ---- ##
idsEligAIDS <- which(active == 1 & stage == "chronic2" &
stage.time != 0 & !is.na(ART.status))
prog.rate.AIDS <- ifelse(ART.status[idsEligAIDS] == 1,
Chronic2ToAIDS.Rate *
ART.Progression.Reduction.Rate,
Chronic2ToAIDS.Rate)
idsAIDS <- idsEligAIDS[rbinom(length(idsEligAIDS), 1,
prog.rate.AIDS) == 1]
if (length(idsAIDS) > 0) {
stage[idsAIDS] <- "AIDS"
5 stage.time[idsAIDS] <- 0
}
## Save attributes ##
dat <- set_attr(dat, "stage", stage)
dat <- set_attr(dat, "stage.time", stage.time)
dat <- set_attr(dat, "ART.status", ART.status)
dat <- set_attr(dat, "ART.time", ART.time)
## Save progression flow statistics ##
dat <- set_epi(dat, "chronic1.ART.flow", at,
sum(ART.status[idsChronic1] == 1))
dat <- set_epi(dat, "chronic1.NoART.flow", at,
sum(ART.status[idsChronic1] == 0))
dat <- set_epi(dat, "chronic2.ART.flow", at,
sum(ART.status[idsChronic2] == 1))
dat <- set_epi(dat, "chronic2.NoART.flow", at,
sum(ART.status[idsChronic2] == 0))
dat <- set_epi(dat, "AIDS.ART.flow", at,
sum(ART.status[idsAIDS] == 1))
dat <- set_epi(dat, "AIDS.NoART.flow", at,
sum(ART.status[idsAIDS] == 0))
return(dat)
}- 1
-
At
at == 2(first module call), initialize custom attributes for all initially infected individuals: acute stage, no ART, and time counters at 0. - 2
-
Increment time counters each step. The
!= 0guard in eligibility checks below prevents individuals from transitioning in the same timestep they entered their current state. - 3
- ART treatment and discontinuance: eligible individuals stochastically start or stop ART. Time counters reset to 0 on transition to enforce the one-step delay.
- 4
- ART reduces progression rates by the reduction factor (default 0.5). Each stage transition uses the same pattern: find eligible, compute ART-adjusted rate, draw Bernoulli outcomes.
- 5
-
Stage transitions reset
stage.timeto 0, preventing double-transitions within a single timestep.
Departure Module
Two departure processes operate in parallel. All susceptible and non-AIDS infected individuals depart at the background rate. Individuals with AIDS depart at an elevated rate, which is further reduced by ART.
dfunc <- function(dat, at) {
## Attributes ##
active <- get_attr(dat, "active")
status <- get_attr(dat, "status")
ART.status <- get_attr(dat, "ART.status")
stage <- get_attr(dat, "stage")
stage.time <- get_attr(dat, "stage.time")
exitTime <- get_attr(dat, "exitTime")
## Parameters ##
departure.rate <- get_param(dat, "departure.rate")
ART.Progression.Reduction.Rate <- get_param(dat,
"ART.Progression.Reduction.Rate")
AIDSToDepart.Rate <- get_param(dat, "AIDSToDepart.Rate")
## ---- Standard departure (susceptible + infected non-AIDS) ---- ##
idsEligStandard <- which(active == 1 &
1 (is.na(stage) | stage != "AIDS"))
idsDepart <- idsEligStandard[rbinom(length(idsEligStandard), 1,
departure.rate) == 1]
## ---- AIDS departure on ART (reduced rate) ---- ##
idsEligAIDSART <- which(active == 1 & stage == "AIDS" &
stage.time != 0 & ART.status == 1 &
!is.na(ART.status))
idsDepartAIDSART <- idsEligAIDSART[rbinom(length(idsEligAIDSART), 1,
AIDSToDepart.Rate *
2 ART.Progression.Reduction.Rate) == 1]
## ---- AIDS departure not on ART ---- ##
idsEligAIDSNoART <- which(active == 1 & stage == "AIDS" &
stage.time != 0 & ART.status == 0 &
!is.na(ART.status))
idsDepartAIDSNoART <- idsEligAIDSNoART[rbinom(length(idsEligAIDSNoART), 1,
3 AIDSToDepart.Rate) == 1]
## Combine all departures and deactivate ##
idsDeparted <- c(idsDepart, idsDepartAIDSART, idsDepartAIDSNoART)
if (length(idsDeparted) > 0) {
active[idsDeparted] <- 0
exitTime[idsDeparted] <- at
}
## Save attributes ##
dat <- set_attr(dat, "active", active)
dat <- set_attr(dat, "exitTime", exitTime)
## Save summary statistics ##
dat <- set_epi(dat, "depart.standard.ART.flow", at,
sum(ART.status[idsDepart] == 1, na.rm = TRUE))
dat <- set_epi(dat, "depart.standard.NoART.flow", at,
sum(ART.status[idsDepart] == 0 |
is.na(ART.status[idsDepart])))
dat <- set_epi(dat, "depart.AIDS.ART.flow", at,
length(idsDepartAIDSART))
dat <- set_epi(dat, "depart.AIDS.NoART.flow", at,
length(idsDepartAIDSNoART))
return(dat)
}- 1
-
Standard departures apply to all susceptibles and infected individuals not yet in the AIDS stage.
is.na(stage)captures susceptibles whose stage attribute isNA. - 2
- AIDS patients on ART depart at the elevated AIDS rate, reduced by the ART progression reduction factor (default 0.5x).
- 3
- AIDS patients not on ART depart at the full elevated AIDS rate — about 1/104 per week, corresponding to a mean 2-year survival from AIDS onset without treatment.
Arrival Module
New individuals arrive as susceptible at a rate proportional to current network size. This uses append_core_attr() for EpiModel 2.x-compatible attribute management.
afunc <- function(dat, at) {
## Parameters ##
n <- sum(get_attr(dat, "active") == 1)
a.rate <- get_param(dat, "arrival.rate")
## Arrival process ##
1 nArrivals <- rpois(1, n * a.rate)
if (nArrivals > 0) {
status <- c(get_attr(dat, "status"), rep("s", nArrivals))
stage <- c(get_attr(dat, "stage"), rep(NA, nArrivals))
stage.time <- c(get_attr(dat, "stage.time"), rep(NA, nArrivals))
ART.status <- c(get_attr(dat, "ART.status"), rep(NA, nArrivals))
ART.time <- c(get_attr(dat, "ART.time"), rep(NA, nArrivals))
2 dat <- append_core_attr(dat, at, nArrivals)
dat <- set_attr(dat, "status", status)
dat <- set_attr(dat, "stage", stage)
dat <- set_attr(dat, "stage.time", stage.time)
dat <- set_attr(dat, "ART.status", ART.status)
3 dat <- set_attr(dat, "ART.time", ART.time)
}
## Summary statistics ##
dat <- set_epi(dat, "a.flow", at, nArrivals)
return(dat)
}- 1
- Number of arrivals is Poisson-distributed with rate proportional to current population size.
- 2
-
append_core_attr()initializes required node attributes (active status, unique ID, entry/exit times). - 3
-
New arrivals are susceptible (
status = "s") with all disease-specific attributes set toNA.
Network Model
Estimation
n <- 1000
1nw <- network_initialize(n)
formation <- ~edges
mean_degree <- 0.8
2target.stats <- c(mean_degree * (n / 2))
departure_rate <- 0.003
coef.diss <- dissolution_coefs(dissolution = ~offset(edges),
duration = 52,
3 d.rate = departure_rate)
coef.diss
est <- netest(nw, formation, target.stats, coef.diss)- 1
- Initialize a network of 1000 nodes representing a sexually active population.
- 2
- Edges-only formation model (Bernoulli random graph). Mean degree of 0.8 means each person has on average 0.8 ongoing partnerships, giving 400 target edges.
- 3
-
Average partnership duration of 52 weeks (1 year). The
d.rateargument adjusts the dissolution coefficient so observed mean duration matches the target after accounting for competing risks from departures.
Dissolution Coefficients
=======================
Dissolution Model: ~offset(edges)
Target Statistics: 52
Crude Coefficient: 3.931826
Mortality/Exit Rate: 0.003
Adjusted Coefficient: 4.305112Network Diagnostics
dx <- netdx(est, nsims = nsims, ncores = ncores, nsteps = nsteps)
Network Diagnostics
-----------------------
- Simulating 5 networks
- Calculating formation statisticsprint(dx)EpiModel Network Diagnostics
=======================
Diagnostic Method: Dynamic
Simulations: 5
Time Steps per Sim: 500
Formation Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 400 398.75 -0.312 2.643 -0.473 7.823 17.522
Duration Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 52 52.073 0.141 0.458 0.16 1.26 2.623
Dissolution Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.019 0.019 1.177 0 1.689 0 0.007plot(dx)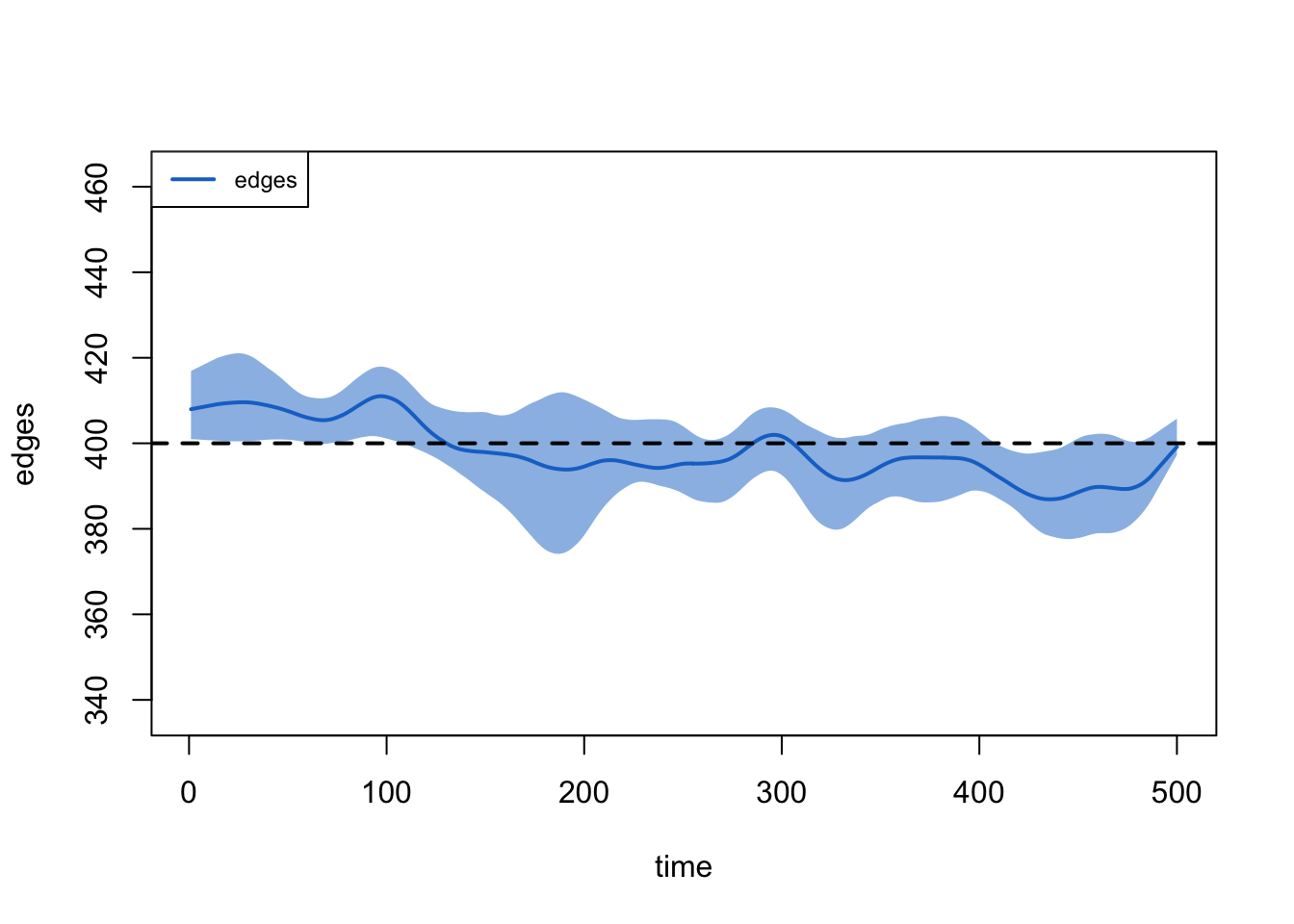
Epidemic Simulation
Scenario 1: With ART
The first scenario models a population where infected individuals can stochastically initiate ART at 1% per week and discontinue at 0.5% per week. ART reduces infectiousness by 95% and halves progression and AIDS mortality rates.
param_art <- param.net(
inf.prob.chronic = 0.01,
relative.inf.prob.acute = 10,
relative.inf.prob.AIDS = 5,
relative.inf.prob.ART = 0.05,
act.rate = 4,
AcuteToChronic1.Rate = 1 / 12,
Chronic1ToChronic2.Rate = 1 / 260,
Chronic2ToAIDS.Rate = 1 / 260,
AIDSToDepart.Rate = 1 / 104,
ART.Treatment.Rate = 0.01,
ART.Discontinuance.Rate = 0.005,
ART.Progression.Reduction.Rate = 0.5,
arrival.rate = 0.002,
departure.rate = departure_rate
)
1init <- init.net(i.num = round(0.05 * n))
control <- control.net(
2 type = NULL,
nsteps = nsteps,
nsims = nsims,
ncores = ncores,
infection.FUN = infect,
progress.FUN = progress,
departures.FUN = dfunc,
arrivals.FUN = afunc,
3 resimulate.network = TRUE,
verbose = TRUE,
module.order = c("resim_nets.FUN",
"progress.FUN",
"infection.FUN",
"departures.FUN",
"arrivals.FUN",
4 "prevalence.FUN")
)
sim_art <- netsim(est, param_art, init, control)
print(sim_art)- 1
- Initial conditions: 5% prevalence (50 of 1000 individuals infected), all initially in the acute stage.
- 2
-
type = NULLmeans the entire module set is custom — no built-in SIS/SIR/SI transmission logic. - 3
- Network resimulation is required because vital dynamics change the active node set each step.
- 4
- Progression runs before infection so that disease stage and ART status are current when transmission probabilities are computed.
EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type:
No. simulations: 5
No. time steps: 500
No. NW groups: 1
Fixed Parameters
---------------------------
act.rate = 4
inf.prob.chronic = 0.01
relative.inf.prob.acute = 10
relative.inf.prob.AIDS = 5
relative.inf.prob.ART = 0.05
AcuteToChronic1.Rate = 0.08333333
Chronic1ToChronic2.Rate = 0.003846154
Chronic2ToAIDS.Rate = 0.003846154
AIDSToDepart.Rate = 0.009615385
ART.Treatment.Rate = 0.01
ART.Discontinuance.Rate = 0.005
ART.Progression.Reduction.Rate = 0.5
arrival.rate = 0.002
departure.rate = 0.003
groups = 1
Model Functions
-----------------------
initialize.FUN
resim_nets.FUN
summary_nets.FUN
infection.FUN
departures.FUN
arrivals.FUN
nwupdate.FUN
prevalence.FUN
verbose.FUN
progress.FUN
Model Output
-----------------------
Variables: s.num i.num num acute.ART.treatment.flow
acute.ART.discontinuance.flow chronic1.ART.treatment.flow
chronic1.ART.discont.flow chronic2.ART.treatment.flow
chronic2.ART.discont.flow AIDS.ART.treatment.flow
AIDS.ART.discontinuance.flow chronic1.ART.flow
chronic1.NoART.flow chronic2.ART.flow chronic2.NoART.flow
AIDS.ART.flow AIDS.NoART.flow acute.flow acute.ART.num
acute.NoART.num chronic1.ART.num chronic1.NoART.num
chronic2.ART.num chronic2.NoART.num AIDS.ART.num
AIDS.NoART.num depart.standard.ART.flow
depart.standard.NoART.flow depart.AIDS.ART.flow
depart.AIDS.NoART.flow a.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 400 408.8 2.2 Inf 0 16.574 16.574
Duration Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 52 66.263 27.428 1.672 8.532 0.51 5.983
Dissolution Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.019 0.013 -31.445 0 -66.89 0 0.004Scenario 2: No ART (Baseline)
The counterfactual scenario uses identical parameters except that the ART treatment rate is set to 0. This lets us isolate the impact of ART on prevalence and incidence — the central question of the Granich et al. model.
param_noart <- param.net(
inf.prob.chronic = 0.01,
relative.inf.prob.acute = 10,
relative.inf.prob.AIDS = 5,
relative.inf.prob.ART = 0.05,
act.rate = 4,
AcuteToChronic1.Rate = 1 / 12,
Chronic1ToChronic2.Rate = 1 / 260,
Chronic2ToAIDS.Rate = 1 / 260,
AIDSToDepart.Rate = 1 / 104,
ART.Treatment.Rate = 0,
ART.Discontinuance.Rate = 0,
ART.Progression.Reduction.Rate = 0.5,
arrival.rate = 0.002,
departure.rate = departure_rate
)
sim_noart <- netsim(est, param_noart, init, control)
print(sim_noart)EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type:
No. simulations: 5
No. time steps: 500
No. NW groups: 1
Fixed Parameters
---------------------------
act.rate = 4
inf.prob.chronic = 0.01
relative.inf.prob.acute = 10
relative.inf.prob.AIDS = 5
relative.inf.prob.ART = 0.05
AcuteToChronic1.Rate = 0.08333333
Chronic1ToChronic2.Rate = 0.003846154
Chronic2ToAIDS.Rate = 0.003846154
AIDSToDepart.Rate = 0.009615385
ART.Treatment.Rate = 0
ART.Discontinuance.Rate = 0
ART.Progression.Reduction.Rate = 0.5
arrival.rate = 0.002
departure.rate = 0.003
groups = 1
Model Functions
-----------------------
initialize.FUN
resim_nets.FUN
summary_nets.FUN
infection.FUN
departures.FUN
arrivals.FUN
nwupdate.FUN
prevalence.FUN
verbose.FUN
progress.FUN
Model Output
-----------------------
Variables: s.num i.num num acute.ART.treatment.flow
acute.ART.discontinuance.flow chronic1.ART.treatment.flow
chronic1.ART.discont.flow chronic2.ART.treatment.flow
chronic2.ART.discont.flow AIDS.ART.treatment.flow
AIDS.ART.discontinuance.flow chronic1.ART.flow
chronic1.NoART.flow chronic2.ART.flow chronic2.NoART.flow
AIDS.ART.flow AIDS.NoART.flow acute.flow acute.ART.num
acute.NoART.num chronic1.ART.num chronic1.NoART.num
chronic2.ART.num chronic2.NoART.num AIDS.ART.num
AIDS.NoART.num depart.standard.ART.flow
depart.standard.NoART.flow depart.AIDS.ART.flow
depart.AIDS.NoART.flow a.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 400 412 3 Inf 0 19.962 19.962
Duration Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 52 65.192 25.368 1.67 7.901 0.691 5.533
Dissolution Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.019 0.013 -30.593 0 -71.553 0 0.004Derived Measures
Before plotting, compute prevalence, incidence rate, ART coverage, and total stage counts by combining ART and non-ART sub-states.
sim_art <- mutate_epi(sim_art,
prev = i.num / num,
ir.rate = acute.flow / s.num,
ART.num = acute.ART.num + chronic1.ART.num +
chronic2.ART.num + AIDS.ART.num
)
sim_art <- mutate_epi(sim_art,
ART.prev = ART.num / i.num
)
sim_noart <- mutate_epi(sim_noart,
prev = i.num / num,
ir.rate = acute.flow / s.num
)
sim_art <- mutate_epi(sim_art,
acute.num = acute.ART.num + acute.NoART.num,
chronic1.num = chronic1.ART.num + chronic1.NoART.num,
chronic2.num = chronic2.ART.num + chronic2.NoART.num,
AIDS.num = AIDS.ART.num + AIDS.NoART.num
)
sim_noart <- mutate_epi(sim_noart,
acute.num = acute.ART.num + acute.NoART.num,
chronic1.num = chronic1.ART.num + chronic1.NoART.num,
chronic2.num = chronic2.ART.num + chronic2.NoART.num,
AIDS.num = AIDS.ART.num + AIDS.NoART.num
)Analysis
HIV Prevalence Comparison
The central question: does ART reduce population-level prevalence? By reducing infectiousness, ART should slow transmission and reduce the equilibrium prevalence over time.
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_noart, y = "prev",
main = "HIV Prevalence: ART vs. No ART",
ylab = "Prevalence", xlab = "Weeks",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim_art, y = "prev", add = TRUE,
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
legend("topleft", legend = c("No ART", "With ART"),
col = c("firebrick", "steelblue"), lwd = 2, bty = "n")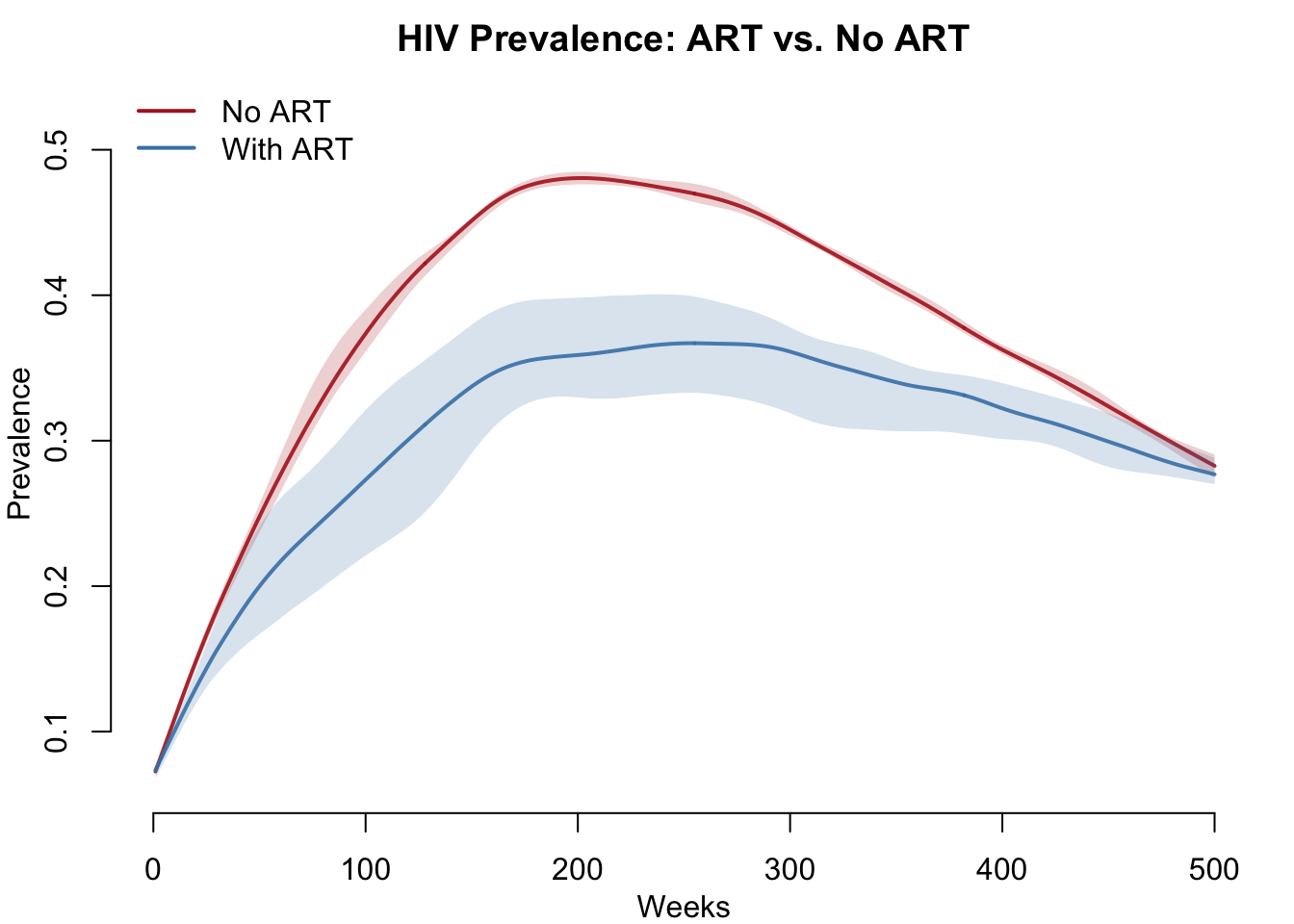
HIV Incidence Rate Comparison
Incidence rate measures new infections per susceptible per week. Because ART reduces the infectiousness of treated individuals, the force of infection acting on susceptibles should be lower in the ART scenario.
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_noart, y = "ir.rate",
main = "HIV Incidence Rate: ART vs. No ART",
ylab = "Incidence Rate", xlab = "Weeks",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim_art, y = "ir.rate", add = TRUE,
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
legend("topleft", legend = c("No ART", "With ART"),
col = c("firebrick", "steelblue"), lwd = 2, bty = "n")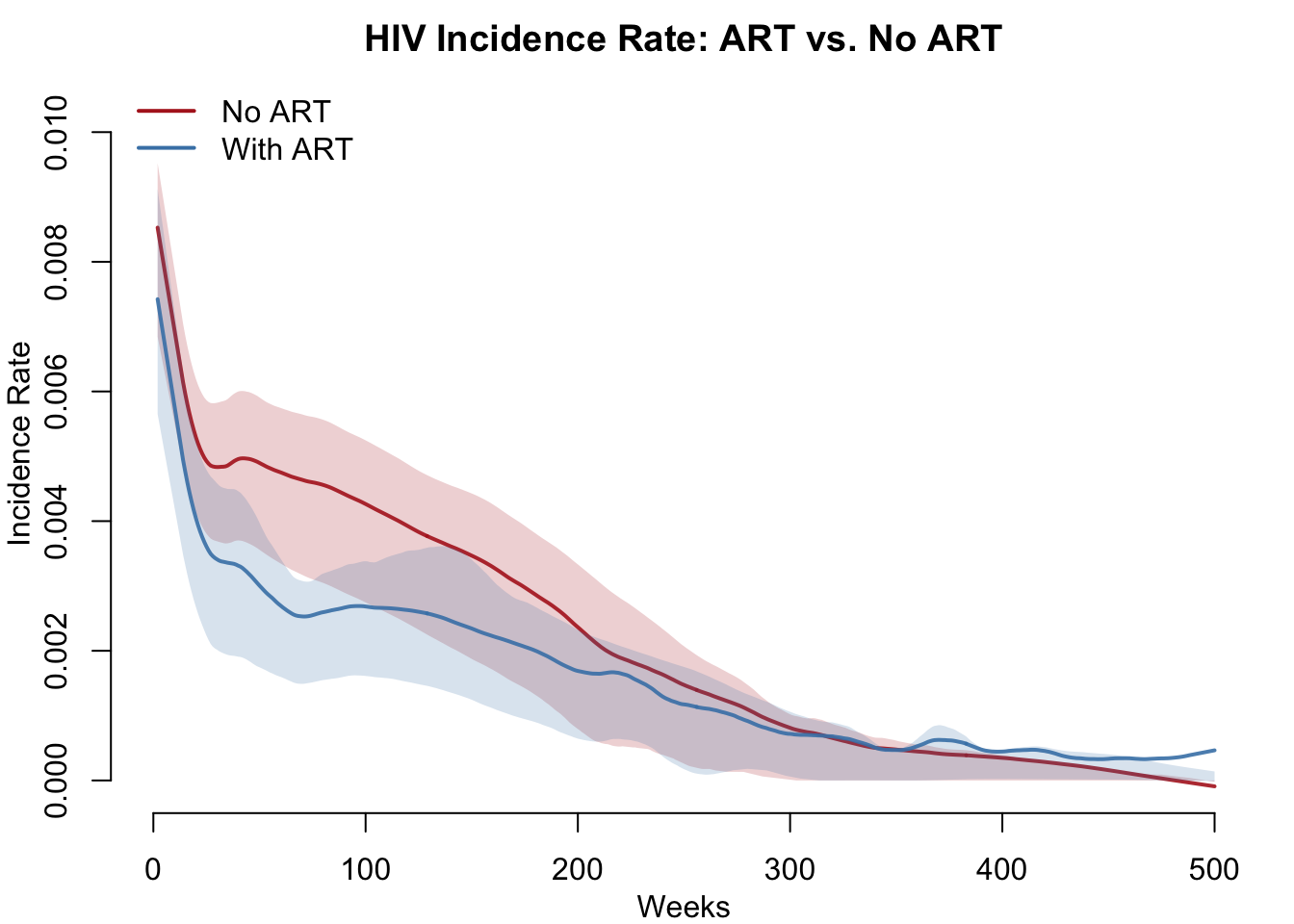
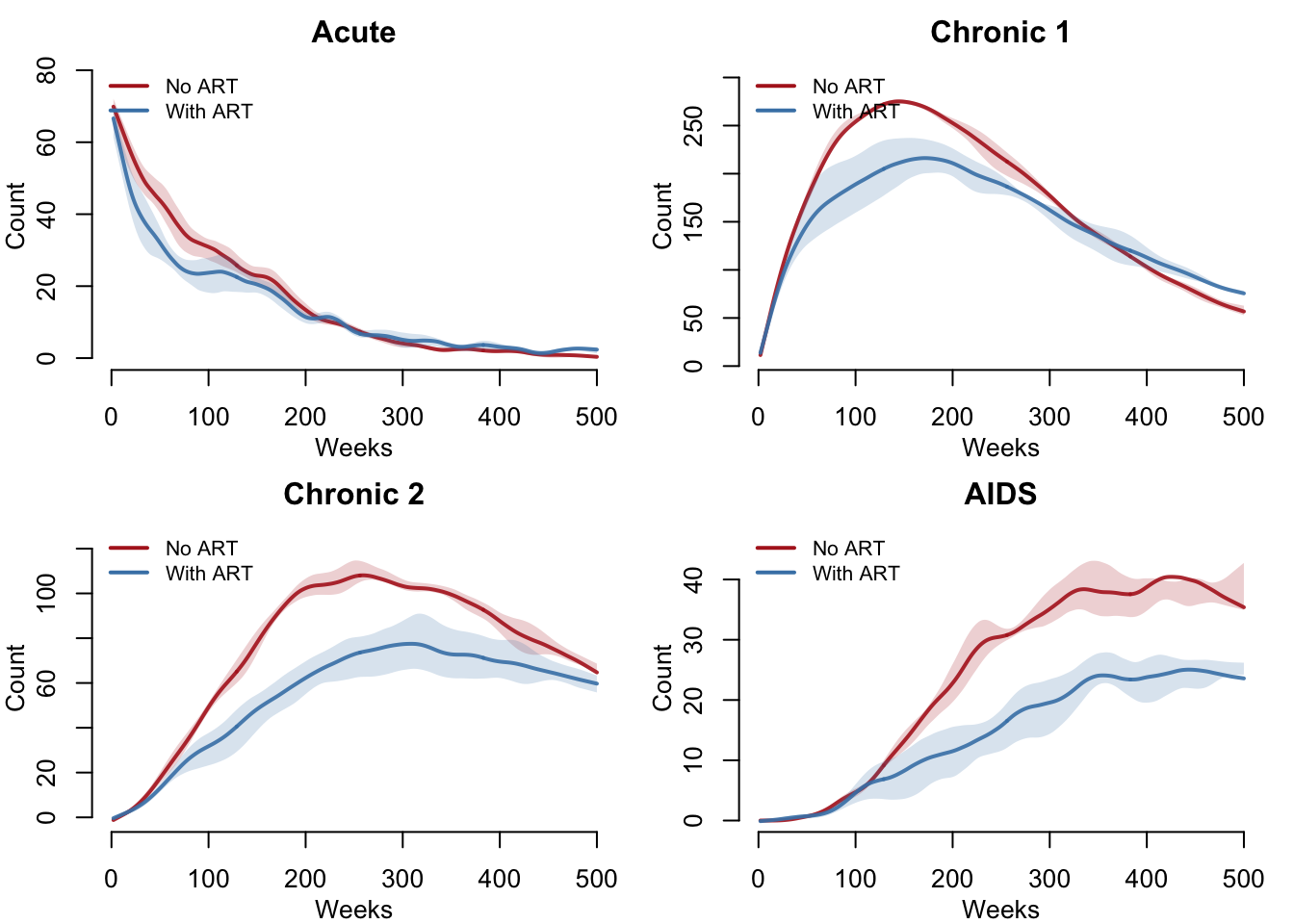
Disease Stage Counts by Scenario
One panel per disease stage, each comparing ART (blue) versus No ART (red). With ART slowing progression, we expect fewer people reaching AIDS and more accumulating in earlier stages.
par(mfrow = c(2, 2), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
stages <- c("acute.num", "chronic1.num", "chronic2.num", "AIDS.num")
titles <- c("Acute", "Chronic 1", "Chronic 2", "AIDS")
for (i in seq_along(stages)) {
plot(sim_noart, y = stages[i],
main = titles[i], ylab = "Count", xlab = "Weeks",
mean.col = "firebrick", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "firebrick", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
plot(sim_art, y = stages[i], add = TRUE,
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
legend = FALSE)
legend("topleft", legend = c("No ART", "With ART"),
col = c("firebrick", "steelblue"), lwd = 2, cex = 0.8, bty = "n")
}
ART Coverage Among PLHIV
What fraction of people living with HIV are on ART? This tracks toward the UNAIDS treatment coverage targets.
par(mfrow = c(1, 1), mar = c(3, 3, 2, 1), mgp = c(2, 1, 0))
plot(sim_art, y = "ART.prev",
main = "ART Coverage Among PLHIV",
ylab = "Proportion on ART", xlab = "Weeks",
mean.col = "steelblue", mean.lwd = 2, mean.smooth = TRUE,
qnts.col = "steelblue", qnts.alpha = 0.2, qnts.smooth = TRUE,
ylim = c(0, 1))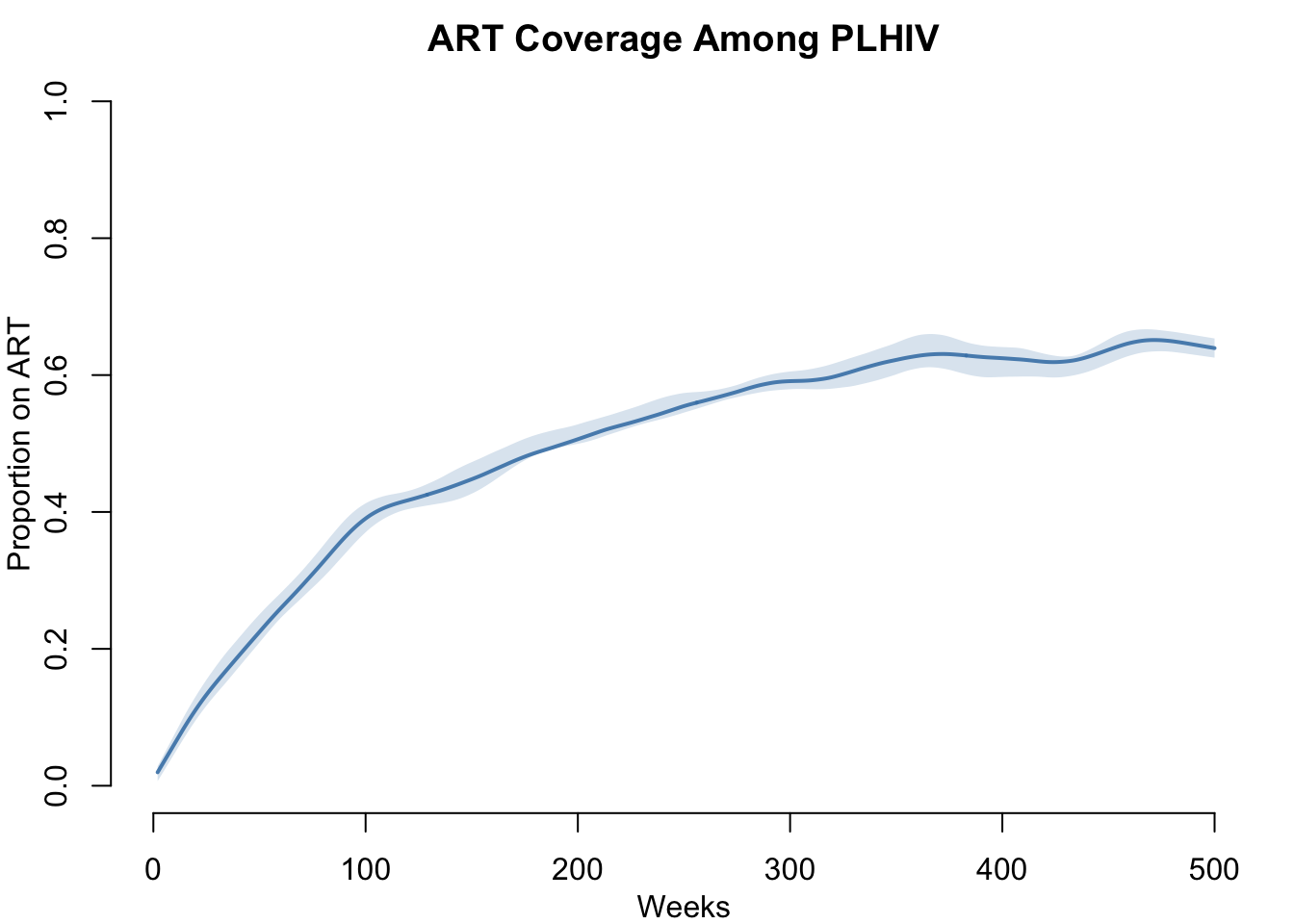
Summary Table
df_art <- as.data.frame(sim_art)
df_noart <- as.data.frame(sim_noart)
last_t <- max(df_art$time)
cum_inf_art <- sum(df_art$acute.flow, na.rm = TRUE)
cum_inf_noart <- sum(df_noart$acute.flow, na.rm = TRUE)
knitr::kable(data.frame(
Metric = c("Cumulative new infections",
"Infections averted by ART",
"Mean prevalence",
"Peak prevalence",
"ART coverage at end (among PLHIV)"),
With_ART = c(cum_inf_art,
cum_inf_noart - cum_inf_art,
round(mean(df_art$prev, na.rm = TRUE), 3),
round(max(df_art$prev, na.rm = TRUE), 3),
round(mean(df_art$ART.prev[df_art$time == last_t]), 3)),
No_ART = c(cum_inf_noart,
NA,
round(mean(df_noart$prev, na.rm = TRUE), 3),
round(max(df_noart$prev, na.rm = TRUE), 3),
NA)
))| Metric | With_ART | No_ART |
|---|---|---|
| Cumulative new infections | 2516.000 | 3221.000 |
| Infections averted by ART | 705.000 | NA |
| Mean prevalence | 0.319 | 0.397 |
| Peak prevalence | 0.412 | 0.539 |
| ART coverage at end (among PLHIV) | 0.634 | NA |
Parameters
Transmission
| Parameter | Description | Default |
|---|---|---|
inf.prob.chronic |
Per-act transmission probability for chronic-stage HIV | 0.01 |
relative.inf.prob.acute |
Multiplier for acute-stage infectiousness | 10 |
relative.inf.prob.AIDS |
Multiplier for AIDS-stage infectiousness | 5 |
relative.inf.prob.ART |
Multiplier for infectiousness while on ART | 0.05 |
act.rate |
Number of acts per partnership per timestep | 4 |
Disease Progression
| Parameter | Description | Default |
|---|---|---|
AcuteToChronic1.Rate |
Per-timestep probability of acute to chronic 1 | 1/12 (~12 weeks) |
Chronic1ToChronic2.Rate |
Per-timestep probability of chronic 1 to chronic 2 | 1/260 (~5 years) |
Chronic2ToAIDS.Rate |
Per-timestep probability of chronic 2 to AIDS | 1/260 (~5 years) |
ART Treatment
| Parameter | Description | Default |
|---|---|---|
ART.Treatment.Rate |
Per-timestep probability of starting ART | 0.01 |
ART.Discontinuance.Rate |
Per-timestep probability of stopping ART | 0.005 |
ART.Progression.Reduction.Rate |
Multiplier applied to progression and AIDS departure rates while on ART | 0.5 |
Vital Dynamics
| Parameter | Description | Default |
|---|---|---|
arrival.rate |
Per-capita arrival rate per timestep | 0.002 |
departure.rate |
Background per-capita departure rate per timestep | 0.003 |
AIDSToDepart.Rate |
Per-timestep departure probability for persons with AIDS | 1/104 (~2 years) |
Network
| Parameter | Description | Default |
|---|---|---|
| Population size | Number of nodes | 1000 |
| Mean degree | Average concurrent partnerships per node | 0.8 |
| Partnership duration | Mean edge duration (weeks) | 52 (~1 year) |
Module Execution Order
resim_nets -> progress -> infect -> departures -> arrivals -> prevalenceProgression runs before infection so that disease stage and ART status are updated before transmission probabilities are computed. This ensures that individuals who just started ART have their reduced infectiousness reflected in the current timestep’s transmission calculations. Departures run after infection to remove individuals from the population, and arrivals follow to replenish the susceptible pool.
Next Steps
- Add a care cascade with testing, linkage to care, and retention stages — see the Syphilis model for a multi-stage diagnosis and treatment pattern
- Model PrEP as a prevention intervention alongside ART-as-prevention, using a similar binary attribute approach
- Add demographic heterogeneity (age, sex, or risk group) with differential network formation — see SI with Vital Dynamics for the age-structured pattern
- Calibrate to empirical data using observed HIV prevalence and ART coverage to set transmission and treatment parameters
- Explore cost-effectiveness by extending with costs and QALYs — see the Cost-Effectiveness Analysis example