library(EpiModel)20 SIS Epidemic Across a Dynamic Network
EpiModel uses separable-temporal exponential-family random graph models (STERGMs) to estimate and simulate complete networks based on individual-level, dyad-level, and network-level patterns of density, degree, assortivity, and other features influencing edge formation and dissolution. Building and simulating network-based epidemic models in EpiModel is a multi-step process, starting with estimation of a temporal ERGM and continuing with simulation of a dynamic network and epidemic processes on top of that network.
In this tutorial, we work through a model of a Susceptible-Infected-Susceptible (SIS) epidemic. One example of an SIS disease would be a bacterial sexually transmitted infection such as Gonorrhea, in which persons may acquire infection from sexual contact with an infected partner, and then recover from infection either through natural clearance or through antibiotic treatment.
We will use a simplifying assumption of a closed population, in which there are no entries or exits from the network; this may be justified by the short time span over which the epidemic will be simulated.
To get started, load the EpiModel library.
20.1 Network Model Estimation
The first step in our network model is to specify a network structure, including features like size and nodal attributes. The network_initialize function creates an object of class network. Below we show an example of initializing a network of 500 nodes, with no edges between them at the start. Edges represent sexual partnerships (mutual person-to-person contact), so this is an undirected network.
nw <- network_initialize(n = 500)The sizes of the networks represented in this workshop are smaller than what might be used for a research-level model, mostly for computational efficiency. Larger network sizes over longer time intervals are typically used for research purposes.
20.1.1 Model Parameterization
This example will start simple, with a formula that represents the network density and the level of concurrency (overlapping sexual partnerships) in the population. This is a dyad-dependent ERGM, since the probability of edge formation between any two nodes depends on the existence of edges between those nodes and other nodes. The concurrent term is defined as the number of nodes with at least two partners at any time. Following the notation of the tergm package, we specify this using a right-hand side (RHS) formula. In addition to concurrency, we will use a constraint on the degree distribution. This will cap the degree of any person at 3, with no nodes allowed to have 4 or more ongoing partnerships. This type of constraint could reflect a truncated sampling scheme for partnerships within a survey (e.g., persons only asked about their 3 most recent partners), or a model assumption about limits of human activity.
formation <- ~edges + concurrent + degrange(from = 4)Target statistics will be the input mechanism for formation model terms. The edges term will be a function of mean degree, or the average number of ongoing partnerships. With an arbitrarily specified mean degree of 0.7, the corresponding target statistic is 175: \(edges = mean \ degree \times \frac{N}{2}\).
We will also specify that 22% of persons exhibit concurrency (this is slightly higher than the 16% expected in a Poisson model conditional on that mean degree). The target statistic for the number of persons with a momentary degree of 4 or more is 0, reflecting our assumed constraint.
target.stats <- c(175, 110, 0)The dissolution model is parameterized from a mean partnership duration estimated from cross-sectional egocentric data. Dissolution models differ from formation models in two respects. First, the dissolution models are not estimated in an ERGM but instead passed in as a fixed coefficient conditional on which the formation model is to be estimated. The dissolution model terms are calculated analytically using the dissolution_coefs function, the output of which is passed into the netest model estimation function. Second, whereas formation models may be arbitrarily complex, dissolution models are limited to a set of dyad-independent models; these are listed in the dissolution_coefs function help page. The model we will use is an edges-only model, implying a homogeneous probability of dissolution for all partnerships in the network. The average duration of these partnerships will be specified at 50 time steps, which will be days in our model.
coef.diss <- dissolution_coefs(dissolution = ~offset(edges), duration = 50)
coef.dissDissolution Coefficients
=======================
Dissolution Model: ~offset(edges)
Target Statistics: 50
Crude Coefficient: 3.89182
Mortality/Exit Rate: 0
Adjusted Coefficient: 3.89182The output from this function indicates both an adjusted and crude coefficient. In this case, they are equivalent. Upcoming workshop material will showcase when they differ as result of exits from the network.
20.1.2 Model Estimation and Diagnostics
In EpiModel, network model estimation is performed with the netest function, which is a wrapper around the estimation functions in the ergm and tergm packages. The function arguments are as follows:
function (nw, formation, target.stats, coef.diss, constraints,
coef.form = NULL, edapprox = TRUE, set.control.ergm = control.ergm(),
set.control.stergm = control.stergm(), set.control.tergm = control.tergm(),
set.control.ergm.ego = control.ergm.ego(), verbose = FALSE,
nested.edapprox = TRUE, ...)
NULLThe four arguments that must be specified with each function call are:
nw: an initialized empty network.formation: a RHS formation formula..target.stats: target statistics for the formation model.coef.diss: output object fromdissolution_coefs, containing the dissolution coefficients.
Other arguments that may be helpful to understand when getting started are:
constraints: this is another way of inputting model constraints (seehelp("ergm")).coef.form: sets the coefficient values of any offset terms in the formation model (those that are not explicitly estimated but fixed).edapprox: selects the dynamic estimation method. IfTRUE, uses the direct method, otherwise the approximation method.- Direct method: uses the functionality of the
tergmpackage to estimate the separable formation and dissolution models for the network. This is often not used because of computational time. - Approximation method: uses
ergmestimation for a cross-sectional network (the prevalence of edges) with an analytic adjustment of the edges coefficient to account for dissolution (i.e., transformation from prevalence to incidence). This approximation method may introduce bias into estimation in certain cases (high density and short durations) but these are typically not a concern for the low density cases in epidemiologically relevant networks.
- Direct method: uses the functionality of the
20.1.2.1 Estimation
Because we have a dyad-dependent model, MCMC will be used to estimate the coefficients of the model given the target statistics.
est <- netest(nw, formation, target.stats, coef.diss)Warning: 'glpk' selected as the solver, but package 'Rglpk' is not available;
falling back to 'lpSolveAPI'. This should be fine unless the sample size and/or
the number of parameters is very big.20.1.2.2 Diagnostics
There are two forms of model diagnostics for a dynamic ERGM fit with netest: static and dynamic diagnostics. When the approximation method has been used, static diagnostics check the fit of the cross-sectional model to target statistics. Dynamic diagnostics check the fit of the model adjusted to account for edge dissolution.
When running a dynamic network simulation, it is good to start with the dynamic diagnostics, and if there are fit problems, work back to the static diagnostics to determine if the problem is due to the cross-sectional fit itself or with the dynamic adjustment (i.e., the approximation method). A proper fitting ERGM using the approximation method does not guarantee well-performing dynamic simulations.
Here we will examine dynamic diagnostics only. These are run with the netdx function, which simulates from the model fit object returned by netest. One must specify the number of simulations from the dynamic model and the number of time steps per simulation. Choice of both simulation parameters depends on the stochasticity in the model, which is a function of network size, model complexity, and other factors. The nwstats.formula contains the network statistics to monitor in the diagnostics: it may contain statistics in the formation model and also others. By default, it is the formation model. Finally, we are keeping the “timed edgelist” with keep.tedgelist.
dx <- netdx(est, nsims = 10, nsteps = 1000,
nwstats.formula = ~edges + meandeg + degree(0:4) + concurrent,
keep.tedgelist = TRUE)We have also built in parallelization into the EpiModel simulation functions, so it is also possible to run multiple simulations at the same time using your computer’s multi-core design. You can find the number of cores in your system with:
parallel::detectCores()Then you can run the multi-core simulations by specifying ncores (EpiModel will prevent you from specifying more cores than you have available).
dx <- netdx(est, nsims = 10, nsteps = 1000, ncores = 4,
nwstats.formula = ~edges + meandeg + degree(0:4) + concurrent,
keep.tedgelist = TRUE)Printing the object will show the object structure and diagnostics. Both formation and duration diagnostics show a good fit relative to their targets. For the formation diagnostics, the mean statistics are the mean of the cross sectional statistics at each time step across all simulations. The Pct Diff column shows the relative difference between the mean and targets. There are two forms of dissolution diagnostics. The edge duration row shows the mean duration of partnerships across the simulations; it tends to be lower than the target unless the diagnostic simulation interval is very long since its average includes a burn-in period where all edges start at a duration of zero (illustrated below in the plot). The next row shows the percent of current edges dissolving at each time step, and is not subject to bias related to burn-in. The percentage of edges dissolution is the inverse of the expected duration: if the duration is 50 days, then we expect that 1/50 (or 2%) to dissolve each day.
print(dx)EpiModel Network Diagnostics
=======================
Diagnostic Method: Dynamic
Simulations: 10
Time Steps per Sim: 1000
Formation Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 175 175.112 0.064 1.446 0.078 5.974 15.079
meandeg NA 0.700 NA 0.006 NA 0.024 0.060
degree0 NA 272.165 NA 1.633 NA 6.095 15.830
degree1 NA 118.734 NA 0.487 NA 1.452 9.468
degree2 NA 95.813 NA 0.700 NA 4.491 11.678
degree3 NA 13.288 NA 0.240 NA 0.837 4.139
degree4 NA 0.000 NA NaN NA 0.000 0.000
concurrent 110 109.101 -0.817 0.936 -0.960 5.183 13.388
Duration Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 50 49.413 -1.174 0.338 -1.739 1.19 3.731
Dissolution Diagnostics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.02 0.02 0.402 0 0.775 0.001 0.011Plotting the diagnostics object will show the time series of the target statistics against any targets. The other options used here specify to smooth the mean lines, give them a thicker line width, and plot each statistic in a separate panel. The black dashed lines show the value of the target statistics for any terms in the model. Similar to the numeric summaries, the plots show a good fit over the time series.
plot(dx)
The simulated network statistics from diagnostic object may be extracted into a data.frame with get_nwstats.
nwstats1 <- get_nwstats(dx, sim = 1)
head(nwstats1, 20) time sim edges meandeg degree0 degree1 degree2 degree3 degree4 concurrent
1 1 1 147 0.588 298 116 80 6 0 86
2 2 1 148 0.592 298 115 80 7 0 87
3 3 1 151 0.604 294 117 82 7 0 89
4 4 1 157 0.628 290 114 88 8 0 96
5 5 1 159 0.636 288 115 88 9 0 97
6 6 1 158 0.632 289 114 89 8 0 97
7 7 1 160 0.640 284 120 88 8 0 96
8 8 1 160 0.640 286 121 80 13 0 93
9 9 1 163 0.652 282 124 80 14 0 94
10 10 1 166 0.664 277 127 83 13 0 96
11 11 1 171 0.684 273 125 89 13 0 102
12 12 1 175 0.700 269 125 93 13 0 106
13 13 1 173 0.692 273 123 89 15 0 104
14 14 1 173 0.692 271 125 91 13 0 104
15 15 1 171 0.684 273 123 93 11 0 104
16 16 1 168 0.672 276 123 90 11 0 101
17 17 1 169 0.676 275 124 89 12 0 101
18 18 1 172 0.688 272 124 92 12 0 104
19 19 1 172 0.688 271 125 93 11 0 104
20 20 1 174 0.696 269 127 91 13 0 104The dissolution model fit may also be assessed with plots by specifying either the duration or dissolution type, as defined above. The duration diagnostic is based on the average age of edges at each time step, up to that time step. An imputation algorithm is used for left-censored edges (i.e., those that exist at t1); you can turn off this imputation to see the effects of censoring with duration.imputed = FALSE. Both metrics show a good fit of the dissolution model to the target duration of 50 time steps.
par(mfrow = c(1, 2))
plot(dx, type = "duration")
plot(dx, type = "dissolution")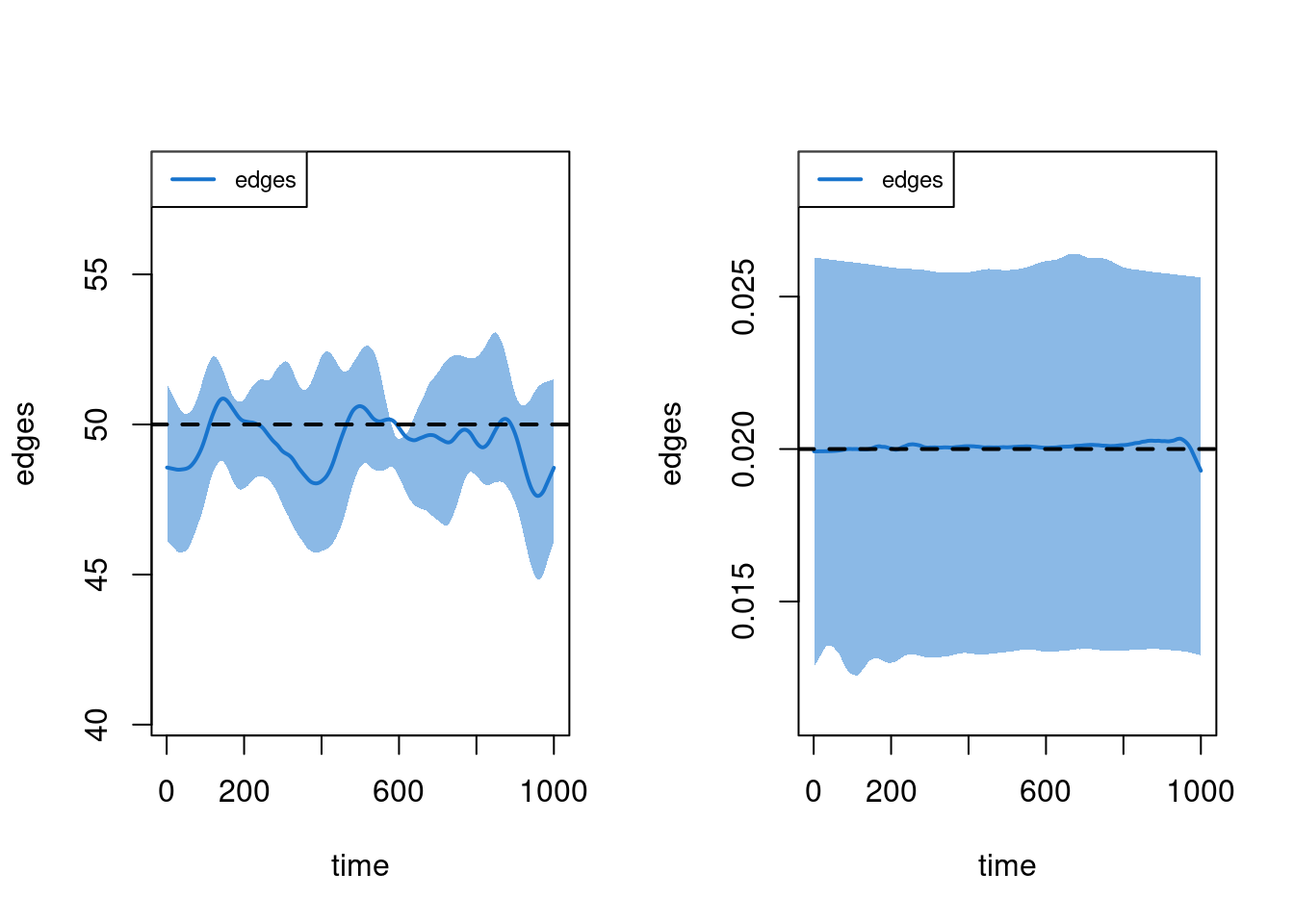
By inspecting the timed edgelist, we can see the burn-in period directly with censoring of onset times. The as.data.frame function is used to extract this edgelist object.
tel <- as.data.frame(dx, sim = 1)
head(tel, 20) onset terminus tail head onset.censored terminus.censored duration edge.id
1 0 54 1 487 TRUE FALSE 54 1
2 0 102 3 132 TRUE FALSE 102 2
3 0 77 7 291 TRUE FALSE 77 3
4 0 129 7 440 TRUE FALSE 129 4
5 0 26 9 99 TRUE FALSE 26 5
6 0 3 13 107 TRUE FALSE 3 6
7 0 67 13 255 TRUE FALSE 67 7
8 0 23 15 48 TRUE FALSE 23 8
9 0 4 18 269 TRUE FALSE 4 9
10 0 25 18 345 TRUE FALSE 25 10
11 0 123 20 392 TRUE FALSE 123 11
12 444 451 20 392 FALSE FALSE 7 11
13 0 36 22 150 TRUE FALSE 36 12
14 0 12 23 91 TRUE FALSE 12 13
15 0 74 30 330 TRUE FALSE 74 14
16 0 41 31 75 TRUE FALSE 41 15
17 0 68 32 158 TRUE FALSE 68 16
18 0 31 33 312 TRUE FALSE 31 17
19 0 38 36 135 TRUE FALSE 38 18
20 301 303 36 135 FALSE FALSE 2 18If the model diagnostics had suggested a poor fit, then additional diagnostics and fitting would be necessary. If using the approximation method, one should first start by running the cross-sectional diagnostics (setting dynamic to FALSE in netdx). Note that the number of simulations may be very large here and there are no time steps specified because each simulation is a cross-sectional network.
dx.static <- netdx(est, nsims = 10000, dynamic = FALSE)
print(dx.static)The plots now represent individual simulations from an MCMC chain, rather than time steps.
par(mfrow = c(1,1))
plot(dx.static, sim.lines = TRUE, sim.lwd = 0.1)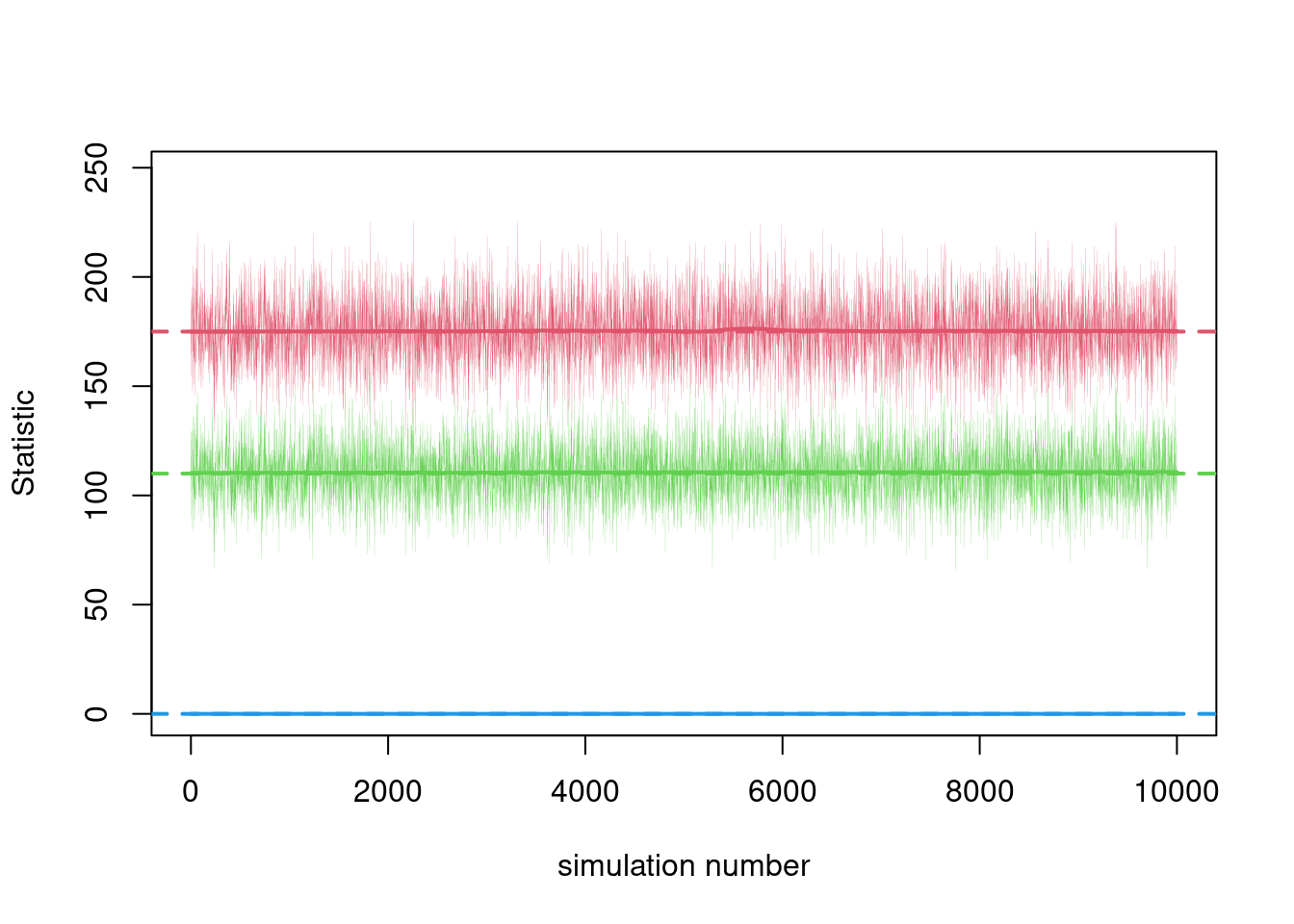
This lack of temporality is now evident when looking at the raw data.
nwstats2 <- get_nwstats(dx.static)
head(nwstats2, 20) sim edges concurrent deg4+
1 1 185 125 0
2 2 165 107 0
3 3 190 129 0
4 4 170 101 0
5 5 176 118 0
6 6 155 90 0
7 7 175 108 0
8 8 180 112 0
9 9 188 116 0
10 10 183 120 0
11 11 196 126 0
12 12 191 130 0
13 13 191 130 0
14 14 185 119 0
15 15 161 101 0
16 16 148 84 0
17 17 152 95 0
18 18 165 102 0
19 19 179 114 0
20 20 191 114 0If the cross-sectional model fits well but the dynamic model does not, then a full STERGM estimation may be necessary (using edapprox = TRUE). If the cross-sectional model does not fit well, different control parameters for the ERGM estimation may be necessary (see the help file for netdx for instructions).
20.2 Epidemic Simulation
EpiModel simulates disease epidemics over dynamic networks by integrating dynamic model simulations with the simulation of other epidemiological processes such as disease transmission and recovery. Like the network model simulations, these processes are also simulated stochastically so that the range of potential outcomes under the model specifications is estimated.
The specification of epidemiological processes to model may be arbitrarily complex, but EpiModel includes a number of “built-in” model types within the software. Additional components will be programmed and plugged into the simulation API (just like any epidemic model); we will start to cover that tomorrow. Here, we will start simple with an SIS epidemic using this built-in functionality. This is starting point to what you can do in EpiModel!
20.2.1 Epidemic Model Parameters
Our SIS model will rely on three parameters. The act rate is the number of sexual acts that occur within a partnership each time unit. The overall frequency of acts per person per unit time is a function of the incidence rate of partnerships and this act rate parameter. The infection probability is the risk of transmission given contact with an infected person. The recovery rate for an SIS epidemic is the speed at which infected persons become susceptible again. For a bacterial STI like gonorrhea, this may be a function of biological attributes like sex or use of therapeutic agents like antibiotics.
EpiModel uses three helper functions to input epidemic parameters, initial conditions, and other control settings for the epidemic model. First, we use the param.net function to input the per-act transmission probability in inf.prob and the number of acts per partnership per unit time in act.rate. The recovery rate implies that the average duration of disease is 10 days (1/rec.rate).
param <- param.net(inf.prob = 0.4, act.rate = 2, rec.rate = 0.1)For initial conditions in this model, we only need to specify the number of infected persons at the outset of the epidemic. The remaining persons in the network will be classified as disease susceptible.
init <- init.net(i.num = 10)The control settings specify the structural elements of the model. These include the disease type, number of simulations, and number of time steps per simulation. (Here again we could use the model multi-core functionality by specifying an ncores value, but these models run so quickly that it’s not necessary.)
control <- control.net(type = "SIS", nsims = 5, nsteps = 500)20.2.2 Simulating the Epidemic Model
Once the model has been parameterized, simulating the model is straightforward. One must pass the fitted network model object from netest along with the parameters, initial conditions, and control settings to the netsim function. With a no-feedback model like this (i.e., there are no vital dynamics parameters), the full dynamic network time series is simulated at the start of each epidemic simulation, and then the epidemiological processes are simulated over that structure.
sim <- netsim(est, param, init, control)Printing the model output lists the inputs and outputs of the model. The output includes the sizes of the compartments (s.num is the number susceptible and i.num is the number infected) and flows (si.flow is the number of infections and is.flow is the number of recoveries). Methods for extracting this output is discussed below.
print(sim)EpiModel Simulation
=======================
Model class: netsim
Simulation Summary
-----------------------
Model type: SIS
No. simulations: 5
No. time steps: 500
No. NW groups: 1
Fixed Parameters
---------------------------
inf.prob = 0.4
act.rate = 2
rec.rate = 0.1
groups = 1
Model Output
-----------------------
Variables: s.num i.num num si.flow is.flow
Networks: sim1 ... sim5
Transmissions: sim1 ... sim5
Formation Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 175 173.795 -0.689 2.254 -0.535 7.888 14.037
concurrent 110 108.386 -1.467 1.615 -0.999 6.785 12.889
deg4+ 0 0.000 NaN NaN NaN 0.000 0.000
Duration Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 50 50.282 0.563 0.562 0.501 2.606 3.959
Dissolution Statistics
-----------------------
Target Sim Mean Pct Diff Sim SE Z Score SD(Sim Means) SD(Statistic)
edges 0.02 0.02 -0.562 0 -0.521 0.001 0.01120.2.3 Model Analysis
Now the the model has been simulated, the next step is to analyze the data. This includes plotting the epidemiological output, the networks over time, and extracting other raw data.
20.2.3.1 Epidemic Plots
Plotting the output from the epidemic model using the default arguments will display the size of the compartments in the model across simulations. The means across simulations at each time step are plotted with lines, and the polygon band shows the inter-quartile range across simulations.
par(mfrow = c(1, 1))
plot(sim)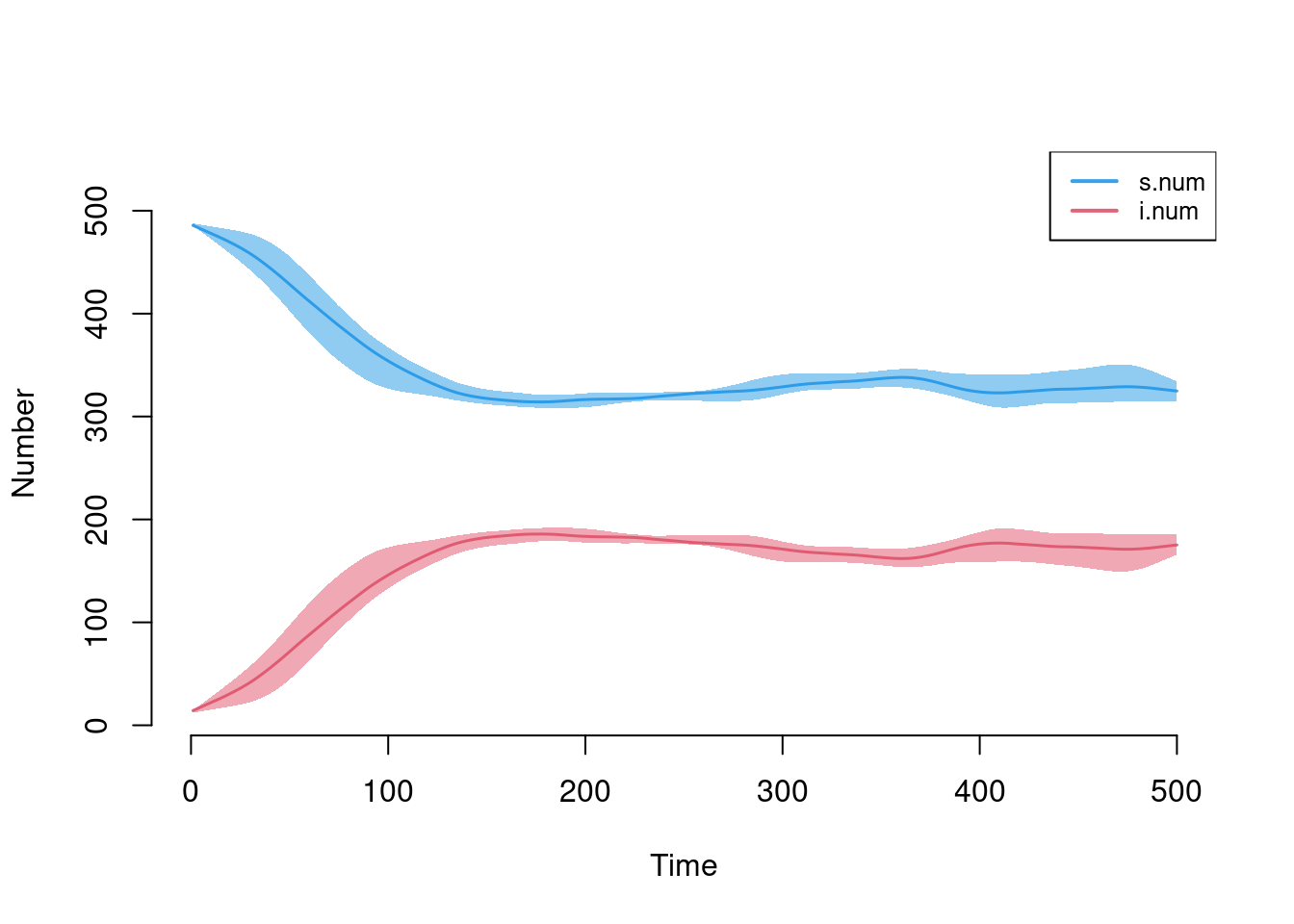
Graphical elements may be toggled on and off. The popfrac argument specifies whether to use the absolute size of compartments versus proportions.
par(mfrow = c(1, 2))
plot(sim, sim.lines = TRUE, mean.line = FALSE, qnts = FALSE, popfrac = TRUE)
plot(sim, mean.smooth = FALSE, qnts = 1, qnts.smooth = FALSE, popfrac = TRUE)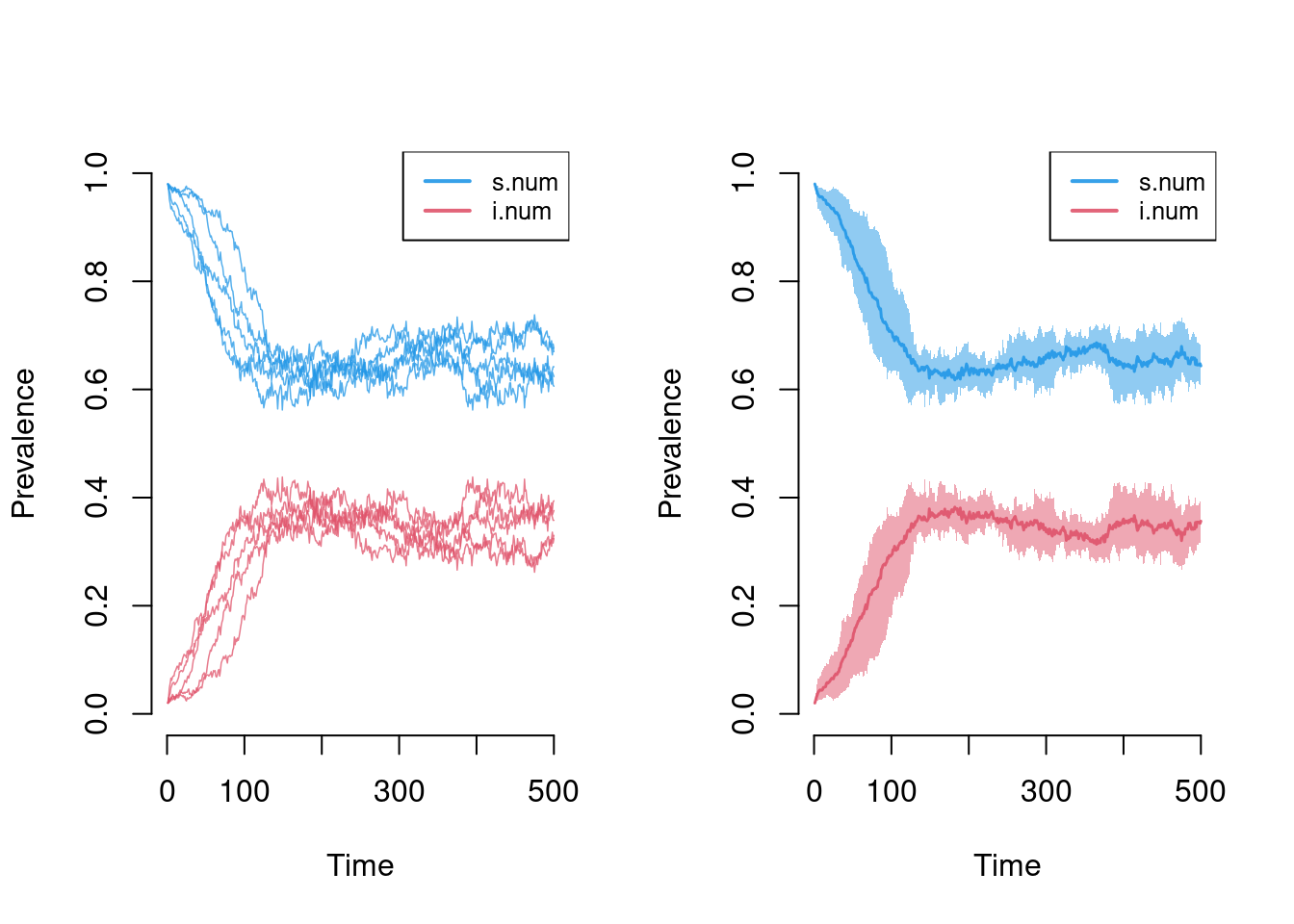
Whereas the default will print the compartment proportions, other elements of the simulation may be plotted by name with the y argument. Here we plot both flow sizes using smoothed means, which converge at model equilibrium by the end of the time series.
par(mfrow = c(1,1))
plot(sim, y = c("si.flow", "is.flow"), qnts = FALSE,
ylim = c(0, 25), legend = TRUE, main = "Flow Sizes")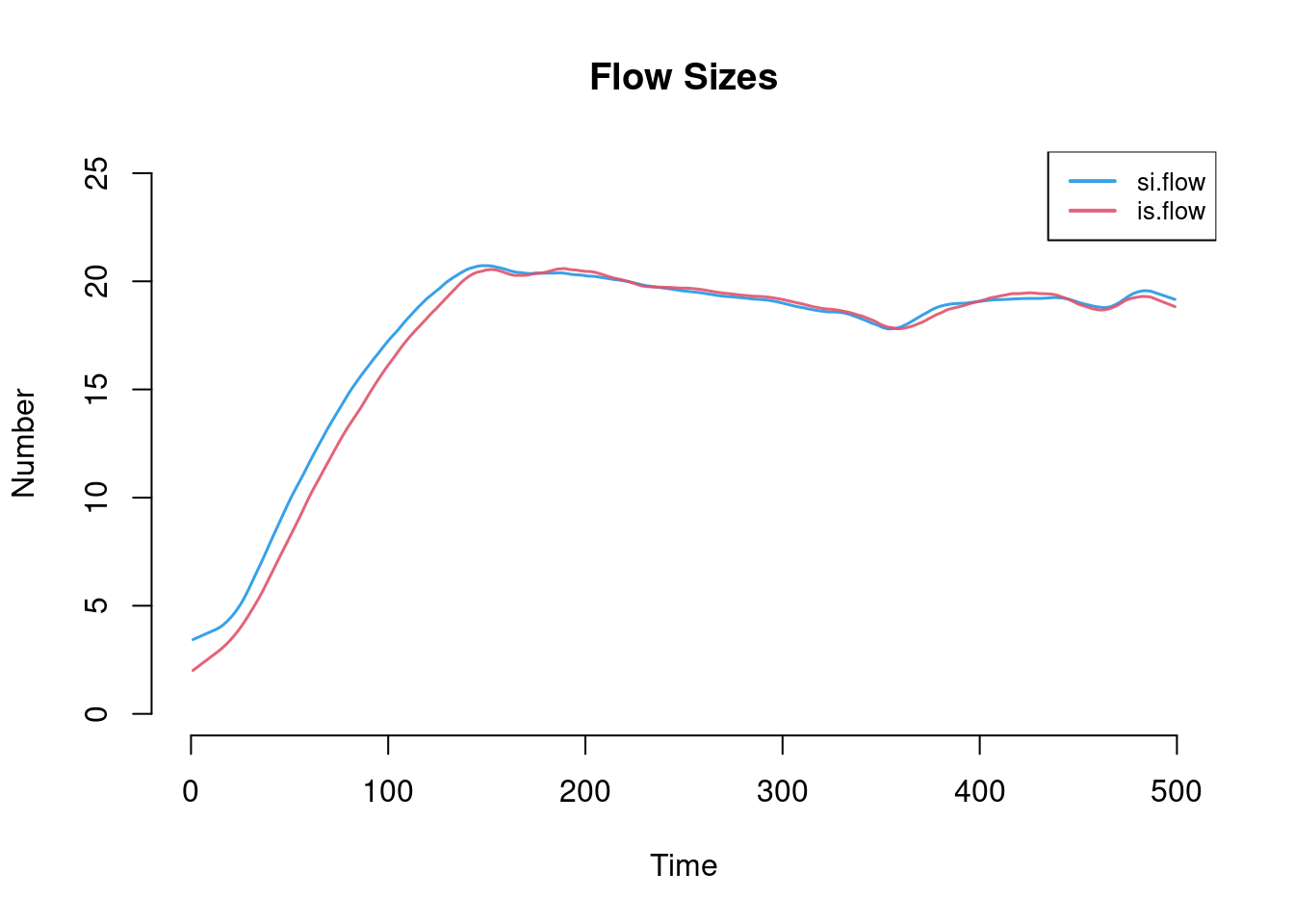
20.2.3.2 Network Plots
Another available plot type is a network plot to visualize the individual nodes and edges at a specific time point. Network plots are output by setting the type parameter to "network". To plot the disease infection status on the nodes, use the col.status argument: blue indicates susceptible and red infected. It is necessary to specify both a time step and a simulation number to plot these networks.
par(mfrow = c(1, 2), mar = c(0, 0, 0, 0))
plot(sim, type = "network", col.status = TRUE, at = 1, sims = 1)
plot(sim, type = "network", col.status = TRUE, at = 500, sims = 1)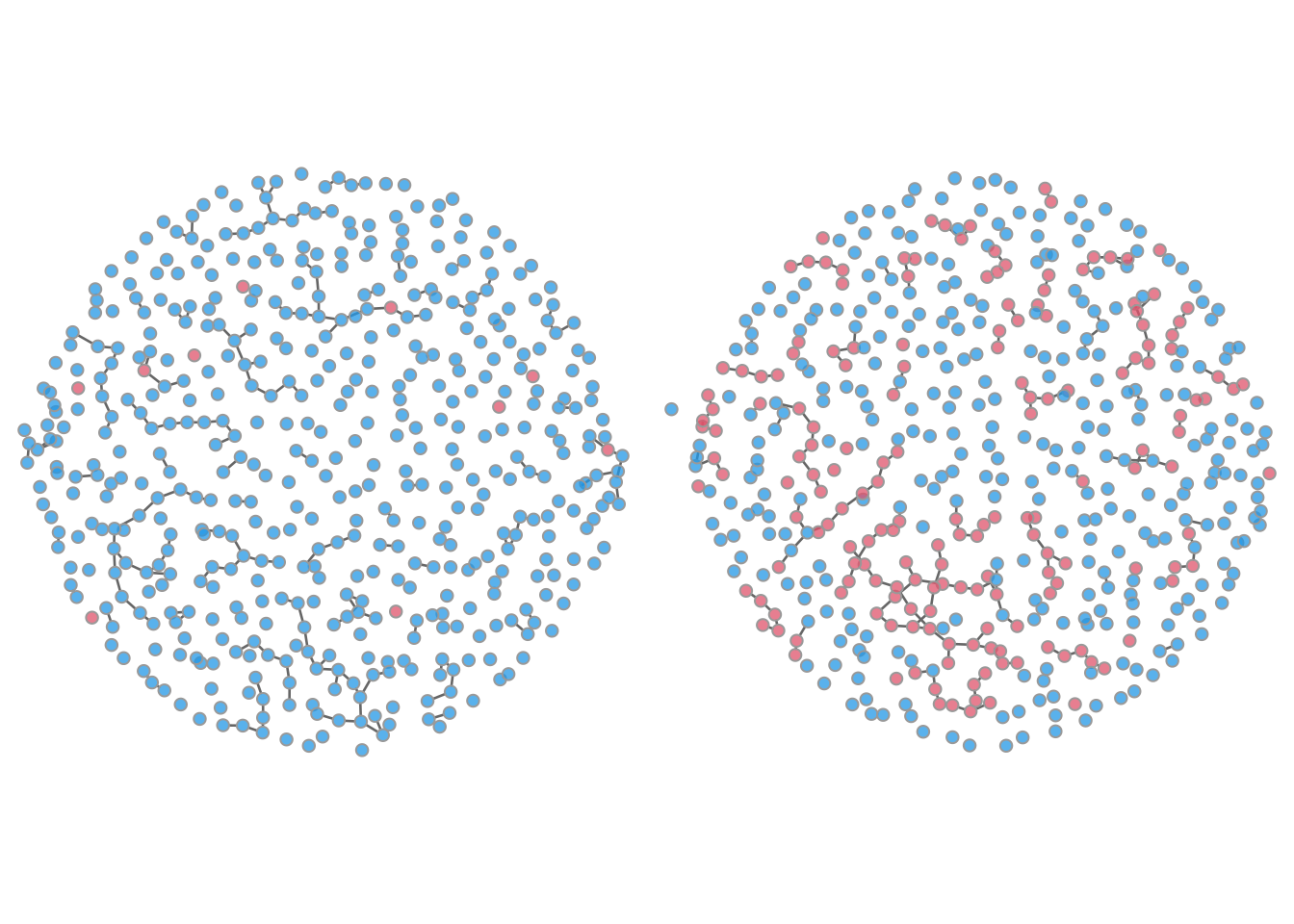
20.2.3.3 Time-Specific Model Summaries
The summary function with the output of netsim will show the model statistics at a specific time step. Here we output the statistics at the final time step, where roughly two-thirds of the population are infected.
summary(sim, at = 500)
EpiModel Summary
=======================
Model class: netsim
Simulation Details
-----------------------
Model type: SIS
No. simulations: 5
No. time steps: 500
No. NW groups: 1
Model Statistics
------------------------------
Time: 500
------------------------------
mean sd pct
Suscept. 321.8 14.890 0.644
Infect. 178.2 14.890 0.356
Total 500.0 0.000 1.000
S -> I 19.2 1.643 NA
I -> S 17.2 2.588 NA
------------------------------ 20.2.3.4 Data Extraction
The as.data.frame function may be used to extract the model output into a data frame object for easy analysis outside of the built-in EpiModel functions. The function default will output the raw data for all simulations for each time step.
df <- as.data.frame(sim)
head(df, 10) sim time s.num i.num num si.flow is.flow
1 1 1 490 10 500 NA NA
2 1 2 489 11 500 4 3
3 1 3 483 17 500 7 1
4 1 4 481 19 500 4 2
5 1 5 476 24 500 7 2
6 1 6 474 26 500 4 2
7 1 7 472 28 500 3 1
8 1 8 471 29 500 3 2
9 1 9 471 29 500 2 2
10 1 10 473 27 500 2 4tail(df, 10) sim time s.num i.num num si.flow is.flow
2491 5 491 321 179 500 23 27
2492 5 492 331 169 500 20 30
2493 5 493 319 181 500 29 17
2494 5 494 321 179 500 19 21
2495 5 495 315 185 500 18 12
2496 5 496 311 189 500 23 19
2497 5 497 309 191 500 22 20
2498 5 498 316 184 500 18 25
2499 5 499 314 186 500 22 20
2500 5 500 312 188 500 21 19The out argument may be changed to specify the output of means across the models (with out = "mean"). The output below shows all compartment and flow sizes as integers, reinforcing this as an individual-level model.
df <- as.data.frame(sim, out = "mean")
head(df, 10) time s.num i.num num si.flow is.flow
1 1 490.0 10.0 500 NaN NaN
2 2 487.4 12.6 500 4.0 1.4
3 3 484.6 15.4 500 5.4 2.6
4 4 482.8 17.2 500 4.0 2.2
5 5 480.4 19.6 500 4.0 1.6
6 6 479.4 20.6 500 3.2 2.2
7 7 478.2 21.8 500 3.2 2.0
8 8 477.8 22.2 500 2.8 2.4
9 9 478.2 21.8 500 2.6 3.0
10 10 478.2 21.8 500 2.8 2.8tail(df, 10) time s.num i.num num si.flow is.flow
491 491 328.6 171.4 500 21.6 20.8
492 492 329.0 171.0 500 21.0 21.4
493 493 323.0 177.0 500 20.8 14.8
494 494 327.4 172.6 500 17.2 21.6
495 495 322.6 177.4 500 20.8 16.0
496 496 322.8 177.2 500 19.8 20.0
497 497 322.6 177.4 500 17.2 17.0
498 498 323.2 176.8 500 18.6 19.2
499 499 323.8 176.2 500 18.2 18.8
500 500 321.8 178.2 500 19.2 17.2The networkDynamic objects are stored in the netsim object, and may be extracted with the get_network function. By default the dynamic networks are saved, and contain the full edge history for every node that has existed in the network, along with the disease status history of those nodes.
nw1 <- get_network(sim, sim = 1)
nw1NetworkDynamic properties:
distinct change times: 503
maximal time range: -Inf until Inf
Dynamic (TEA) attributes:
Vertex TEAs: testatus.active
Includes optional net.obs.period attribute:
Network observation period info:
Number of observation spells: 2
Maximal time range observed: 0 until 501
Temporal mode: discrete
Time unit: step
Suggested time increment: 1
Network attributes:
vertices = 500
directed = FALSE
hyper = FALSE
loops = FALSE
multiple = FALSE
bipartite = FALSE
net.obs.period: (not shown)
total edges= 1940
missing edges= 0
non-missing edges= 1940
Vertex attribute names:
active status testatus.active vertex.names
Edge attribute names not shown One thing you can do with that network dynamic object is to extract the timed edgelist of all ties that existed for that simulation.
nwdf <- as.data.frame(nw1)
head(nwdf, 25) onset terminus tail head onset.censored terminus.censored duration edge.id
1 0 116 1 327 TRUE FALSE 116 1
2 0 6 3 146 TRUE FALSE 6 2
3 0 22 3 162 TRUE FALSE 22 3
4 0 10 3 393 TRUE FALSE 10 4
5 0 16 5 53 TRUE FALSE 16 5
6 0 1 5 137 TRUE FALSE 1 6
7 0 24 6 43 TRUE FALSE 24 7
8 0 57 6 372 TRUE FALSE 57 8
9 0 17 7 83 TRUE FALSE 17 9
10 0 33 7 407 TRUE FALSE 33 10
11 0 48 8 145 TRUE FALSE 48 11
12 0 12 9 269 TRUE FALSE 12 12
13 0 46 11 108 TRUE FALSE 46 13
14 0 33 16 314 TRUE FALSE 33 14
15 0 52 18 398 TRUE FALSE 52 15
16 0 7 21 240 TRUE FALSE 7 16
17 0 184 21 479 TRUE FALSE 184 17
18 0 27 22 60 TRUE FALSE 27 18
19 0 272 22 388 TRUE FALSE 272 19
20 0 17 28 359 TRUE FALSE 17 20
21 0 121 31 430 TRUE FALSE 121 21
22 0 16 32 105 TRUE FALSE 16 22
23 0 102 32 117 TRUE FALSE 102 23
24 0 5 32 426 TRUE FALSE 5 24
25 0 2 33 66 TRUE FALSE 2 25A matrix is stored that records some key details about each transmission event that occurred. Shown below are the first 10 transmission events for simulation number 1. The sus column shows the unique ID of the previously susceptible, newly infected node in the event. The inf column shows the ID of the transmitting node. The other columns show the duration of the transmitting node’s infection at the time of transmission, the per-act transmission probability, act rate during the transmission, and final per-partnership transmission rate (which is the per-act probability raised to the number of acts).
tm1 <- get_transmat(sim, sim = 1)
head(tm1, 10)# A tibble: 10 × 7
# Groups: at, sus [10]
at sus inf infDur transProb actRate finalProb
<dbl> <int> <int> <dbl> <dbl> <dbl> <dbl>
1 2 52 316 4 0.4 2 0.64
2 2 404 204 8 0.4 2 0.64
3 2 429 204 8 0.4 2 0.64
4 2 446 194 4 0.4 2 0.64
5 3 100 316 5 0.4 2 0.64
6 3 204 429 1 0.4 2 0.64
7 3 267 404 1 0.4 2 0.64
8 3 371 194 5 0.4 2 0.64
9 3 445 52 1 0.4 2 0.64
10 3 447 95 5 0.4 2 0.6420.2.3.5 Data Exporting and Plotting with ggplot
We built in plotting methods directly for netsim class objects so you can easily plot multiple types of summary statistics from the simulated model object. However, if you prefer an external plotting tool in R, such as ggplot, it is easy to extract the data in tidy format for analysis and plotting. Here is an example how to do so for out model above. See the help for the ggplot if you are unfamiliar with this syntax.
df <- as.data.frame(sim)
df.mean <- as.data.frame(sim, out = "mean")
library(ggplot2)
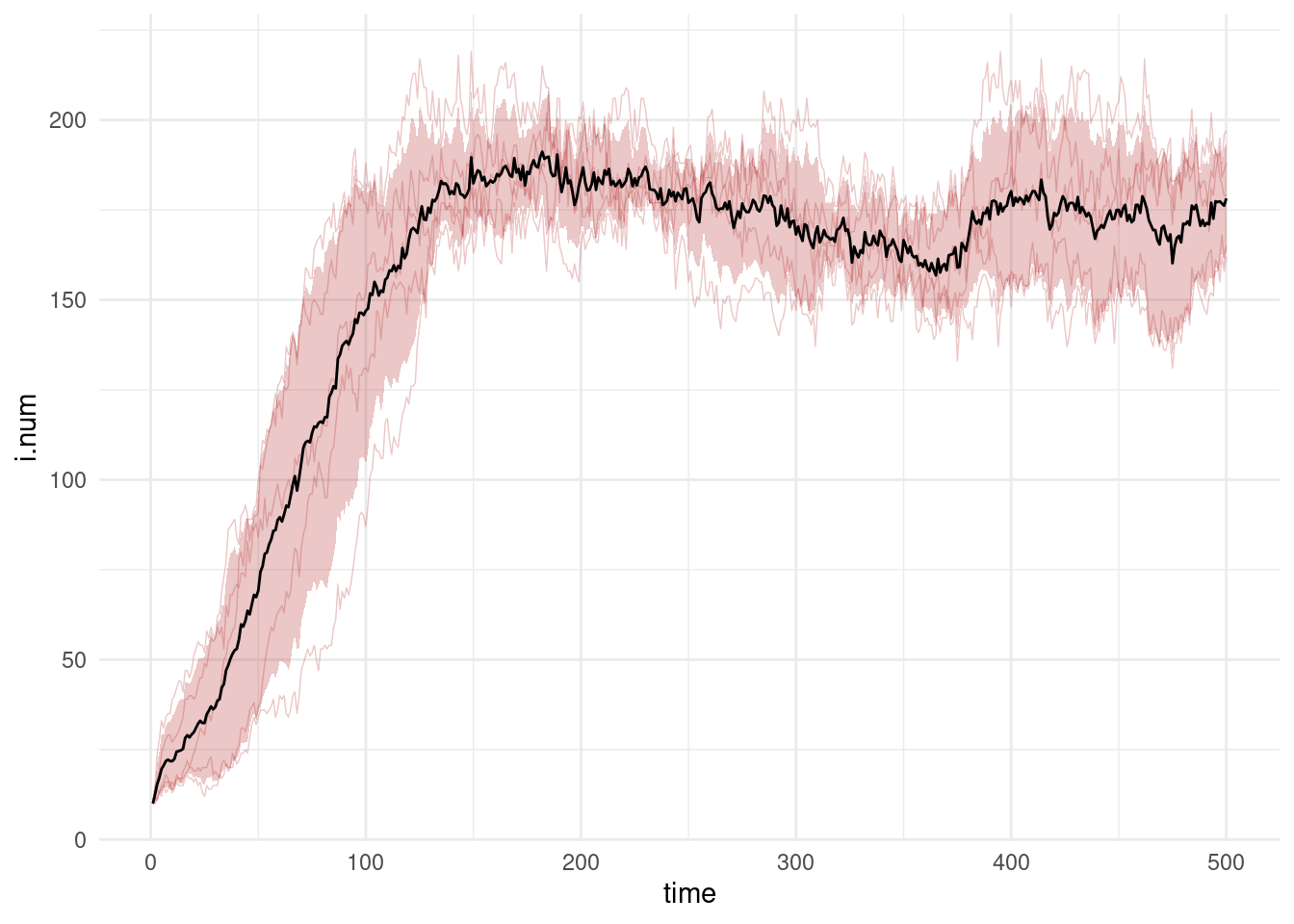
ggplot() +
geom_line(data = df, mapping = aes(time, i.num, group = sim), alpha = 0.25,
lwd = 0.25, color = "firebrick") +
geom_bands(data = df, mapping = aes(time, i.num),
lower = 0.1, upper = 0.9, fill = "firebrick") +
geom_line(data = df.mean, mapping = aes(time, i.num)) +
theme_minimal()